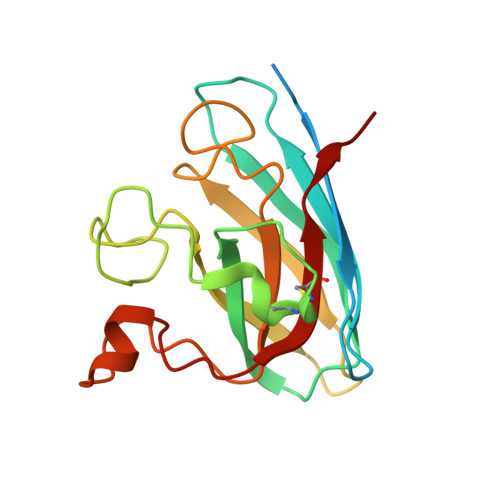
Structural insights into the modulation Of SOD1 aggregation By a fungal metabolite Phialomustin-B: Therapeutic potential in ALS.
Unni, S., Kommu, P., Aouti, S., Nalli, Y., Bharath, M.M.S., Ali, A., Padmanabhan, B.(2024) PLoS One 19: e0298196-e0298196
- PubMed: 38446760 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0298196
- Primary Citation Related Structures:
7XX3 - PubMed Abstract:
Amyotrophic lateral sclerosis (ALS) is a fatal human motor neuron disease leading to muscle atrophy and paralysis. Mutations in superoxide dismutase 1 (SOD1) are associated with familial ALS (fALS). The SOD1 mutants in ALS have a toxic-gain of function by destabilizing the functional SOD1 homodimer, consequently inducing fibril-like aggregation with a cytotoxic non-native trimer intermediate. Therefore, reducing SOD1 oligomerization via chemical modulators is an optimal therapy in ALS. Here, we report the discovery of Phialomustin-B, an unsaturated secondary metabolite from the endophytic fungus Phialophora mustea, as a modulator of SOD1 aggregation. The crystal structure of the SOD1-Phialomustin complex refined to 1.90 Å resolution demonstrated for the first time that the ligand binds to the dimer interface and the lateral region near the electrostatic loop. The aggregation analyses of SOD1WT and the disease mutant SOD1A4V revealed that Phialomustin-B reduces cytotoxic trimerization. We propose that Phialomustin-B is a potent lead molecule with therapeutic potential in fALS.
- Department of Biophysics, National Institute of Mental Health and Neurosciences, Bengaluru, India.
Organizational Affiliation: