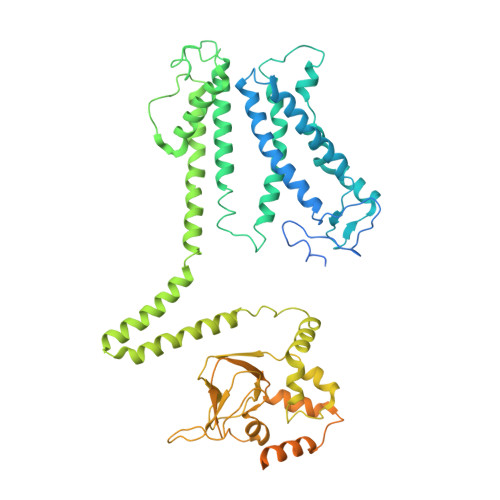
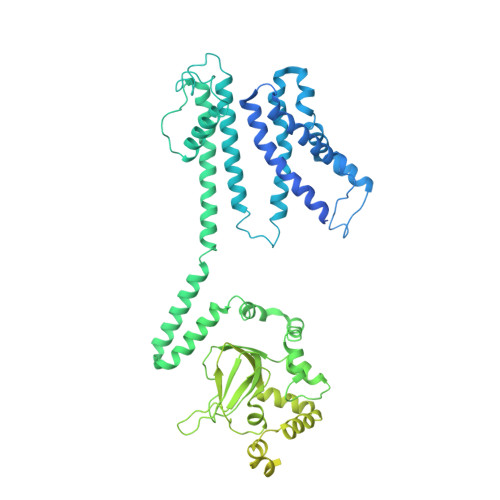
Structural basis for the activity regulation of a potassium channel AKT1 from Arabidopsis.
Lu, Y., Yu, M., Jia, Y., Yang, F., Zhang, Y., Xu, X., Li, X., Yang, F., Lei, J., Wang, Y., Yang, G.(2022) Nat Commun 13: 5682-5682
- PubMed: 36167696 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33420-8
- Primary Citation Related Structures:
7FCV, 7WSW, 7XUF - PubMed Abstract:
The voltage-gated potassium channel AKT1 is responsible for primary K + uptake in Arabidopsis roots. AKT1 is functionally activated through phosphorylation and negatively regulated by a potassium channel α-subunit AtKC1. However, the molecular basis for the modulation mechanism remains unclear. Here we report the structures of AKT1, phosphorylated-AKT1, a constitutively-active variant, and AKT1-AtKC1 complex. AKT1 is assembled in 2-fold symmetry at the cytoplasmic domain. Such organization appears to sterically hinder the reorientation of C-linkers during ion permeation. Phosphorylated-AKT1 adopts an alternate 4-fold symmetric conformation at cytoplasmic domain, which indicates conformational changes associated with symmetry switch during channel activation. To corroborate this finding, we perform structure-guided mutagenesis to disrupt the dimeric interface and identify a constitutively-active variant Asp379Ala mediates K + permeation independently of phosphorylation. This variant predominantly adopts a 4-fold symmetric conformation. Furthermore, the AKT1-AtKC1 complex assembles in 2-fold symmetry. Together, our work reveals structural insight into the regulatory mechanism for AKT1.
- State Key Laboratory for Agrobiotechnology, Frontiers Science Center for Molecular Design Breeding, College of Biological Sciences, China Agricultural University, Beijing, China.
Organizational Affiliation: