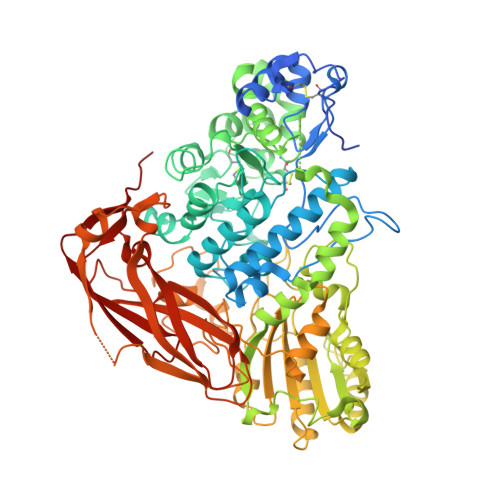
Crystal structure and identification of amino acid residues for catalysis and binding of GH3 AnBX beta-xylosidase from Aspergillus niger.
Kaenying, W., Choengpanya, K., Tagami, T., Wattana-Amorn, P., Lang, W., Okuyama, M., Li, Y.K., Kimura, A., Kongsaeree, P.T.(2023) Appl Microbiol Biotechnol 107: 2335-2349
- PubMed: 36877249 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00253-023-12445-z
- Primary Citation Related Structures:
7XTJ - PubMed Abstract:
β-Xylosidases catalyze the hydrolysis of xylooligosaccharides to xylose in the final step of hemicellulose degradation. AnBX, which is a GH3 β-xylosidase from Aspergillus niger, has a high catalytic efficiency toward xyloside substrates. In this study, we report the three-dimensional structure and the identification of catalytic and substrate binding residues of AnBX by performing site-directed mutagenesis, kinetic analysis, and NMR spectroscopy-associated analysis of the azide rescue reaction. The structure of the E88A mutant of AnBX, determined at 2.5-Å resolution, contains two molecules in the asymmetric unit, each of which is composed of three domains, namely an N-terminal (β/α) 8 TIM-barrel-like domain, an (α/β) 6 sandwich domain, and a C-terminal fibronectin type III domain. Asp288 and Glu500 of AnBX were experimentally confirmed to act as the catalytic nucleophile and acid/base catalyst, respectively. The crystal structure revealed that Trp86, Glu88 and Cys289, which formed a disulfide bond with Cys321, were located at subsite -1. Although the E88D and C289W mutations reduced catalytic efficiency toward all four substrates tested, the substitution of Trp86 with Ala, Asp and Ser increased the substrate preference for glucoside relative to xyloside substrates, indicating that Trp86 is responsible for the xyloside specificity of AnBX. The structural and biochemical information of AnBX obtained in this study provides invaluable insight into modulating the enzymatic properties for the hydrolysis of lignocellulosic biomass. KEY POINTS: • Asp288 and Glu500 of AnBX are the nucleophile and acid/base catalyst, respectively • Glu88 and the Cys289-Cys321 disulfide bond are crucial for the catalytic activity of AnBX • The W86A and W86S mutations in AnBX increased the preference for glucoside substrates.
- Department of Biochemistry, Faculty of Science, Kasetsart University, Bangkok, 10900, Thailand.
Organizational Affiliation: