Crystal structure of CBP bromodomain liganded with Y08175
Xiang, Q., Wang, C., Wu, T., Zhang, Y., Zhang, C., Luo, G., Wu, X., Shen, H., Xu, Y.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Isoform 2 of CREB-binding protein | 136 | Homo sapiens | Mutation(s): 0 Gene Names: CREBBP, CBP EC: 2.3.1.48 (PDB Primary Data), 2.3.1 (PDB Primary Data) | 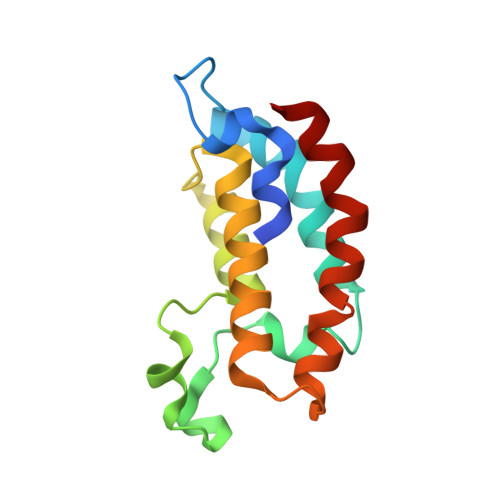 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q92793 GTEx: ENSG00000005339 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q92793 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EJ3 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | B [auth A] | 3-[(1-ethanoyl-5-methoxy-indol-3-yl)carbonylamino]-4-fluoranyl-5-(1-methylpyrazol-4-yl)benzoic acid C23 H19 F N4 O5 RCPJEOKKJODBRQ-UHFFFAOYSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | D [auth A], E [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| DMS Download:Ideal Coordinates CCD File | C [auth A] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 56.89 | α = 90 |
| b = 56.89 | β = 90 |
| c = 80.62 | γ = 120 |
| Software Name | Purpose |
|---|---|
| Aimless | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| iMOSFLM | data reduction |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 22007088 |