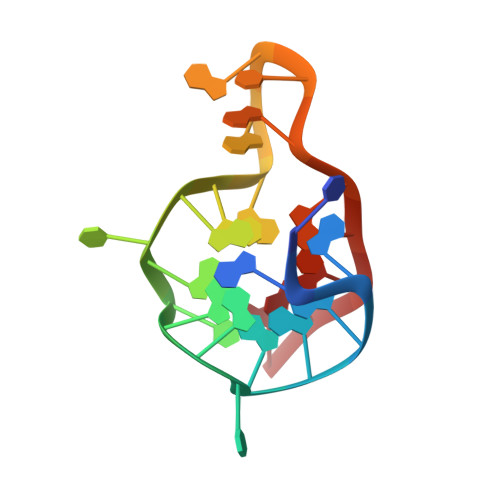
Structural insight into the bulge-containing KRAS oncogene promoter G-quadruplex bound to berberine and coptisine.
Wang, K.B., Liu, Y., Li, J., Xiao, C., Wang, Y., Gu, W., Li, Y., Xia, Y.Z., Yan, T., Yang, M.H., Kong, L.Y.(2022) Nat Commun 13: 6016-6016
- PubMed: 36224201 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33761-4
- Primary Citation Related Structures:
7X8M, 7X8N, 7X8O - PubMed Abstract:
KRAS is one of the most highly mutated oncoproteins, which is overexpressed in various human cancers and implicated in poor survival. The G-quadruplex formed in KRAS oncogene promoter (KRAS-G4) is a transcriptional modulator and amenable to small molecule targeting. However, no available KRAS-G4-ligand complex structure has yet been determined, which seriously hinders the structure-based rational design of KRAS-G4 targeting drugs. In this study, we report the NMR solution structures of a bulge-containing KRAS-G4 bound to berberine and coptisine, respectively. The determined complex structure shows a 2:1 binding stoichiometry with each compound recruiting the adjacent flacking adenine residue to form a "quasi-triad plane" that stacks over the two external G-tetrads. The binding involves both π-stacking and electrostatic interactions. Moreover, berberine and coptisine significantly lowered the KRAS mRNA levels in cancer cells. Our study thus provides molecular details of ligand interactions with KRAS-G4 and is beneficial for the design of specific KRAS-G4-interactive drugs.
- Jiangsu Key Laboratory of Bioactive Natural Product Research and State Key Laboratory of Natural Medicines, Department of Natural Medicinal Chemistry, China Pharmaceutical University, Nanjing, 210009, China. kbwang@cpu.edu.cn.
Organizational Affiliation: