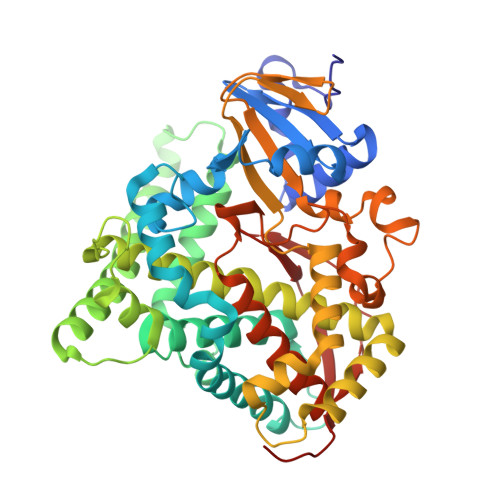
A Compound I Mimic Reveals the Transient Active Species of a Cytochrome P450 Enzyme: Insight into the Stereoselectivity of P450-Catalysed Oxidations.
Suzuki, K., Stanfield, J.K., Omura, K., Shisaka, Y., Ariyasu, S., Kasai, C., Aiba, Y., Sugimoto, H., Shoji, O.(2023) Angew Chem Int Ed Engl 62: e202215706-e202215706
- PubMed: 36519803 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202215706
- Primary Citation Related Structures:
7WY1, 7WY2, 7WY3, 7WY4 - PubMed Abstract:
Catching the structure of cytochrome P450 enzymes in flagrante is crucial for the development of P450 biocatalysts, as most structures collected are found trapped in a precatalytic conformation. At the heart of P450 catalysis lies Cpd I, a short-lived, highly reactive intermediate, whose recalcitrant nature has thwarted most attempts at capturing catalytically relevant poses of P450s. We report the crystal structure of P450BM3 mimicking the state in the precise moment preceding epoxidation, which is in perfect agreement with the experimentally observed stereoselectivity. This structure was attained by incorporation of the stable Cpd I mimic oxomolybdenum mesoporphyrin IX into P450BM3 in the presence of styrene. The orientation of styrene to the Mo-oxo species in the crystal structures sheds light onto the dynamics involved in the rotation of styrene to present its vinyl group to Cpd I. This method serves as a powerful tool for predicting and modelling the stereoselectivity of P450 reactions.
- Department of Chemistry, Graduate School of Science, Nagoya University Furo-cho, Chikusa-ku, Nagoya, 464-8602, Japan.
Organizational Affiliation: