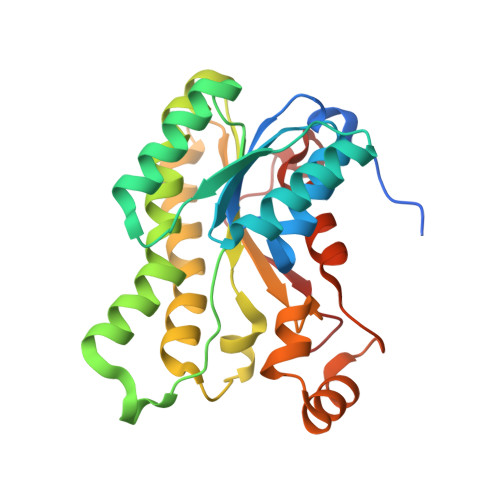
Crystal structure of L-arabinose 1-dehydrogenase as a short-chain reductase/dehydrogenase protein.
Watanabe, S., Yoshiwara, K., Matsubara, R., Watanabe, Y.(2022) Biochem Biophys Res Commun 604: 14-21
- PubMed: 35279441 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2022.03.028
- Primary Citation Related Structures:
7WWX - PubMed Abstract:
l-Arabinose 1-dehydrogenase (AraDH) catalyzes the NAD(P) + -dependent oxidation of l-arabinose to L-arabinono-1,4-lactone in the non-phosphorylative l-arabinose pathway, and is classified into glucose-fructose oxidoreductase and short-chain dehydrogenase/reductase (SDR). We herein report the crystal structure of a SDR-type AraDH (from Herbaspirillum huttiense) for the first time. The interactions between Asp49 and the 2'- and 3'-hydroxyl groups of NAD + were consistent with strict specificity for NAD + . In a binding model for the substrate, Ser155 and Tyr168, highly conserved in the SDR superfamily, interacted with the C1 and/or C2 hydroxyl(s) of l-arabinose, whereas interactions between Asp107, Arg109, and Gln206 and the C2 and/or C3 hydroxyl(s) were unique to AraDH. Trp200 significantly contributed to the selectivities of the C4 hydroxyl and C6 methyl of substrates.
- Faculty of Agriculture, Ehime University, 3-5-7 Tarumi, Matsuyama, Ehime, 790-8566, Japan; Department of Bioscience, Graduate School of Agriculture, Ehime University, 3-5-7 Tarumi, Matsuyama, Ehime, 790-8566, Japan; Center for Marine Environmental Studies (CMES), Ehime University, 2-5 Bunkyo-cho, Matsuyama, Ehime, 790-8577, Japan. Electronic address: irab@agr.ehime-u.ac.jp.
Organizational Affiliation: