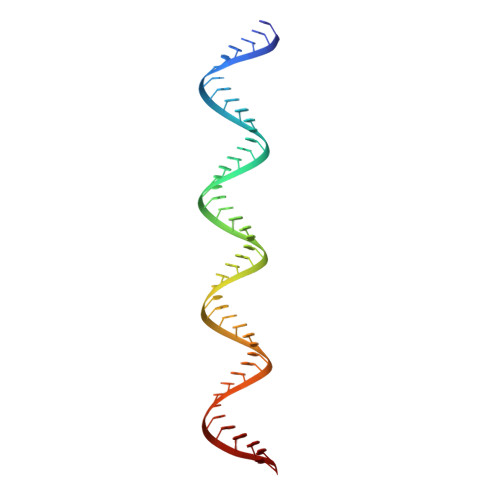
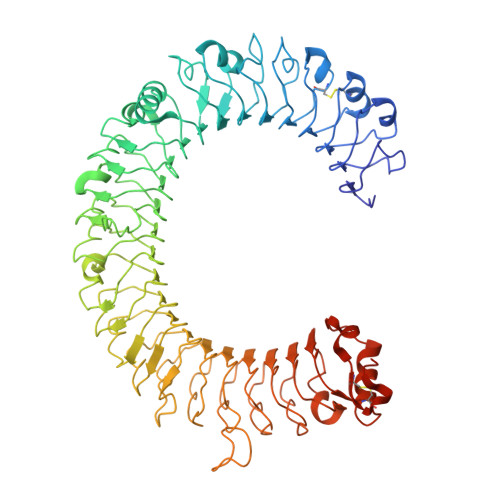
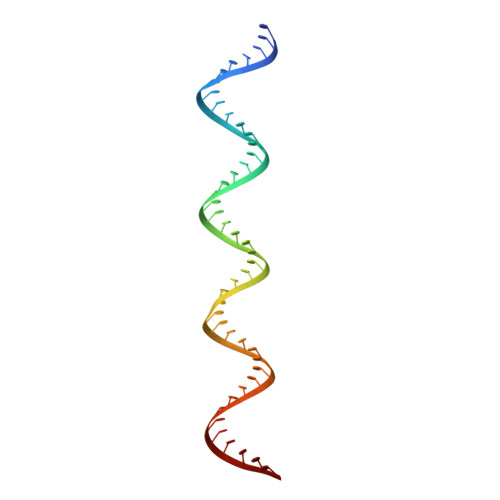TLR3 forms a highly organized cluster when bound to a poly(I:C) RNA ligand.
Lim, C.S., Jang, Y.H., Lee, G.Y., Han, G.M., Jeong, H.J., Kim, J.W., Lee, J.O.(2022) Nat Commun 13: 6876-6876
- PubMed: 36371424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34602-0
- Primary Citation Related Structures:
7WV3, 7WV4, 7WV5, 7WVE, 7WVF, 7WVJ - PubMed Abstract:
Toll-like Receptor 3 (TLR3) initiates a potent anti-viral immune response by binding to double-stranded RNA ligands. Previous crystallographic studies showed that TLR3 forms a homodimer when bound to a 46-base pair RNA ligand. However, this short RNA fails to initiate a robust immune response. To obtain structural insights into the length dependency of TLR3 ligands, we determine the cryo-electron microscopy structure of full-length TLR3 in a complex with a synthetic RNA ligand with an average length of ~400 base pairs. In the structure, the dimeric TLR3 units are clustered along the double-stranded RNA helix in a highly organized and cooperative fashion with a uniform inter-dimer spacing of 103 angstroms. The intracellular and transmembrane domains are dispensable for the clustering because their deletion does not interfere with the cluster formation. Our structural observation suggests that ligand-induced clustering of TLR3 dimers triggers the ordered assembly of intracellular signaling adaptors and initiates a robust innate immune response.
- Department of Life Sciences and POSTECH, Pohang, 37673, Korea.
Organizational Affiliation: