Cryo-EM structure of dodecamer P97 at 2.99 Angstroms resolution
Liu, S., Wang, T.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transitional endoplasmic reticulum ATPase | 755 | Homo sapiens | Mutation(s): 0 Gene Names: VCP EC: 3.6.4.6 | 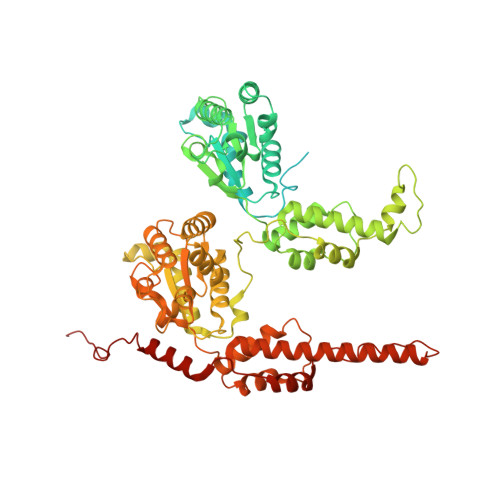 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P55072 GTEx: ENSG00000165280 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55072 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transitional endoplasmic reticulum ATPase | 576 | Homo sapiens | Mutation(s): 0 Gene Names: VCP EC: 3.6.4.6 | 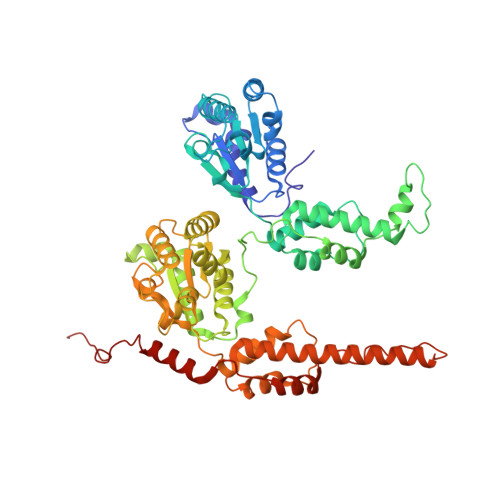 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P55072 GTEx: ENSG00000165280 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55072 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transitional endoplasmic reticulum ATPase | 576 | Homo sapiens | Mutation(s): 0 Gene Names: VCP EC: 3.6.4.6 | 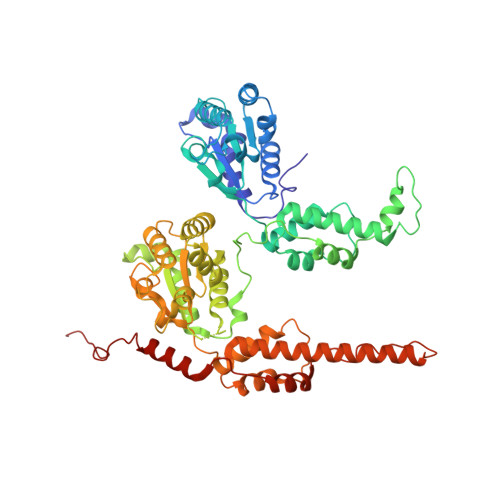 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P55072 GTEx: ENSG00000165280 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55072 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| Y6Y (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | BA [auth H] DA [auth I] FA [auth J] HA [auth K] JA [auth L] | 3-[3-cyclopentylsulfanyl-5-[[3-methyl-4-(4-methylsulfonylphenyl)phenoxy]methyl]-1,2,4-triazol-4-yl]pyridine C27 H28 N4 O3 S2 UJGTUKMAJVCBIS-UHFFFAOYSA-N |  | ||
| ADP Download:Ideal Coordinates CCD File | AA [auth H] CA [auth I] EA [auth J] GA [auth K] IA [auth L] | ADENOSINE-5'-DIPHOSPHATE C10 H15 N5 O10 P2 XTWYTFMLZFPYCI-KQYNXXCUSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | -- |