Cross-species recognition and molecular basis of SARS-CoV-2 and SARS-CoV binding to ACE2s of marine animals.
Li, S., Yang, R., Zhang, D., Han, P., Xu, Z., Chen, Q., Zhao, R., Zhao, X., Qu, X., Zheng, A., Wang, L., Li, L., Hu, Y., Zhang, R., Su, C., Niu, S., Zhang, Y., Qi, J., Liu, K., Wang, Q., Gao, G.F.(2022) Natl Sci Rev 9: nwac122-nwac122
- PubMed: 36187898 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nsr/nwac122
- Primary Citation Related Structures:
7WSE, 7WSF, 7WSG, 7WSH - PubMed Abstract:
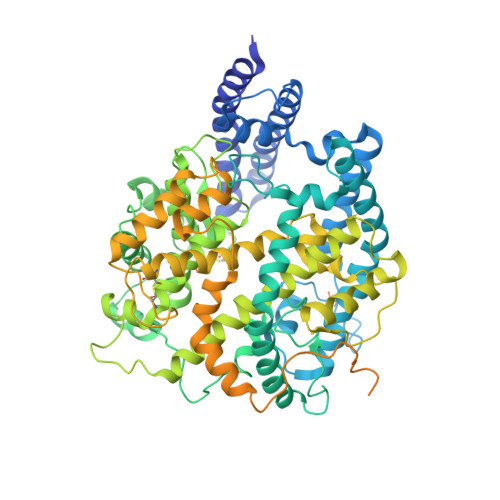
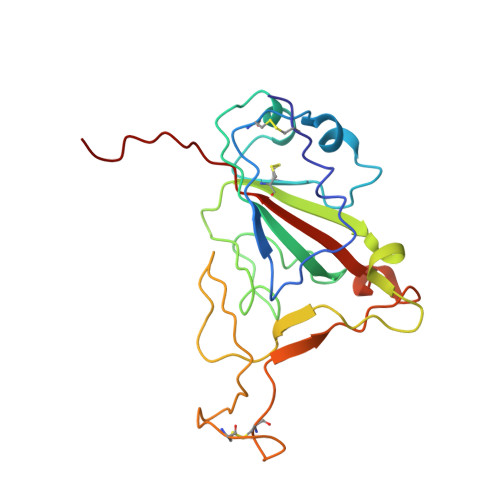
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has an extremely broad host range that includes hippopotami, which are phylogenetically closely related to whales. The cellular ACE2 receptor is one of the key determinants of the host range. Here, we found that ACE2s from several marine mammals and hippopotami could efficiently bind to the receptor-binding domain (RBD) of both SARS-CoV and SARS-CoV-2 and facilitate the transduction of SARS-CoV and SARS-CoV-2 pseudoviruses into ACE2-expressing cells. We further resolved the cryo-electron microscopy complex structures of the minke whale ACE2 and sea lion ACE2, respectively, bound to the RBDs, revealing that they have similar binding modes to human ACE2 when it comes to the SARS-CoV-2 RBD and SARS-CoV RBD. Our results indicate that marine mammals could potentially be new victims or virus carriers of SARS-CoV-2, which deserves further careful investigation and study. It will provide an early warning for the prospective monitoring of marine mammals.
- CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing100101, China.
Organizational Affiliation: