Molecular basis of receptor binding and antibody neutralization of Omicron
Hong, Q., Han, W., Li, J., Xu, S., Wang, Y., Xu, C., Li, Z., Wang, Y., Zhang, C., Huang, Z., Cong, Y.(2022) Nature 604: 546-552
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Angiotensin-converting enzyme 2 | 625 | Homo sapiens | Mutation(s): 0 Gene Names: ACE2, UNQ868/PRO1885 EC: 3.4.17.23 (PDB Primary Data), 3.4.17 (PDB Primary Data) | 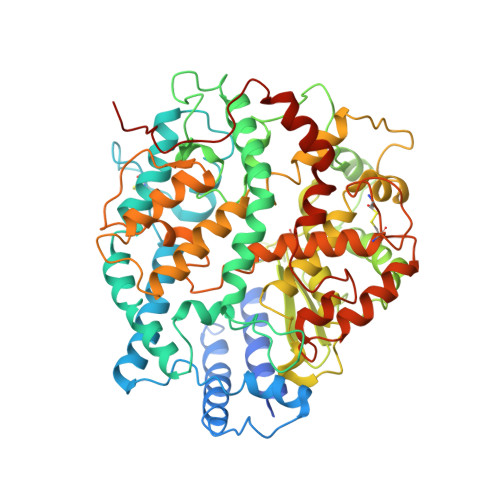 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9BYF1 GTEx: ENSG00000130234 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9BYF1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Spike glycoprotein | 1,258 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 35 Gene Names: S, 2 | 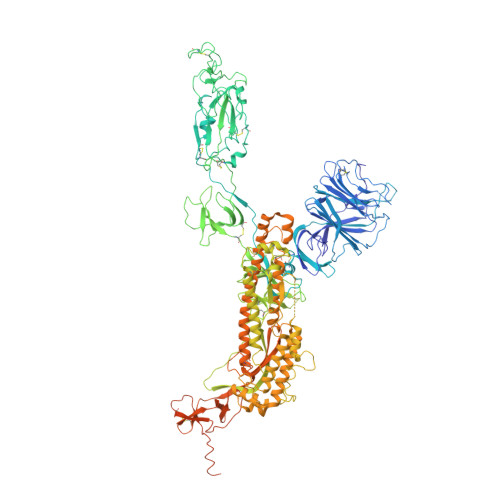 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTC2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Chinese Academy of Sciences | China | -- |