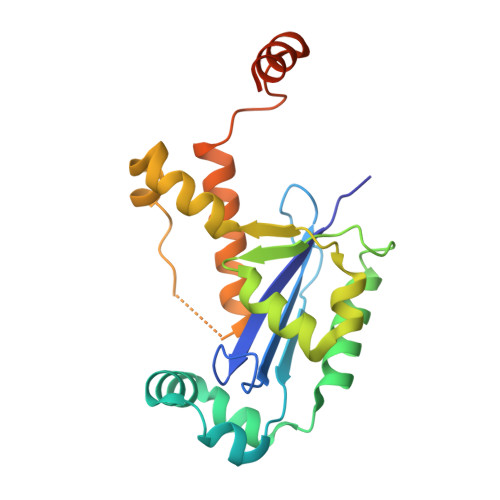
Three-dimensional structure of a mycobacterial oligoribonuclease reveals a unique C-terminal tail that stabilizes the homodimer.
Badhwar, P., Khan, S.H., Taneja, B.(2022) J Biological Chem 298: 102595-102595
- PubMed: 36244449 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102595
- Primary Citation Related Structures:
7VH4, 7WIK - PubMed Abstract:
Oligoribonucleases (Orns) are highly conserved DnaQ-fold 3'-5' exoribonucleases that have been found to carry out the last step of cyclic-di-GMP (c-di-GMP) degradation, that is, pGpG to GMP in several bacteria. Removal of pGpG is critical for c-di-GMP homeostasis, as excess uncleaved pGpG can have feedback inhibition on phosphodiesterases, thereby perturbing cellular signaling pathways regulated by c-di-GMP. Perturbation of c-di-GMP levels not only affects survival under hypoxic, reductive stress, or nutrient-limiting conditions but also affects pathogenicity in infection models as well as antibiotic response in mycobacteria. Here, we have determined the crystal structure of MSMEG_4724, the Orn of Mycobacterium smegmatis (Ms_orn) to 1.87 Å resolution to investigate the function of its extended C-terminal tail that is unique among bacterial Orns. Ms_orn is a homodimer with the canonical RNase-H fold of exoribonucleases and conserved catalytic residues in the active site. Further examination of the substrate-binding site with a modeled pGpG emphasized the role of a phosphate cap and "3'OH cap" in constricting a 2-mer substrate in the active site. The unique C-terminal tail of Ms_orn aids dimerization by forming a handshake-like flap over the second protomer of the dimer. Our thermal and denaturant-induced unfolding experiments suggest that it helps in higher stability of Ms_orn as compared with Escherichia coli Orn or a C-terminal deletion mutant. We also show that the C-terminal tail is required for modulating response to stress agents in vivo. These results will help in further evaluating the role of signaling and regulation by c-di-GMP in mycobacteria.
- CSIR-Institute of Genomics and Integrative Biology (CSIR-IGIB), New Delhi, India; Academy of Scientific and Innovative Research (AcSIR), Ghaziabad, India.
Organizational Affiliation: