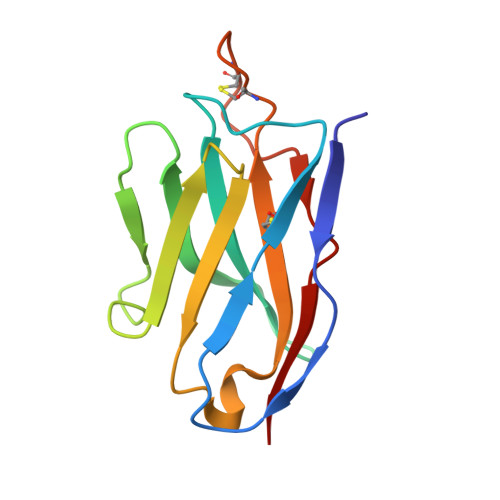
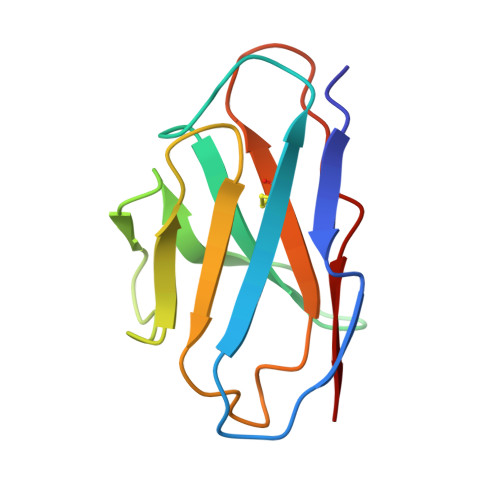
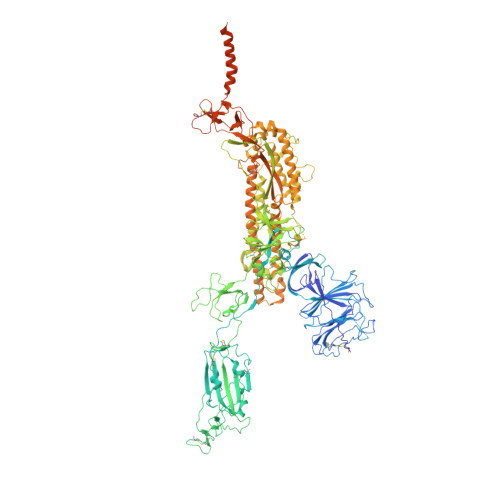
Memory B cell repertoire from triple vaccinees against diverse SARS-CoV-2 variants.
Wang, K., Jia, Z., Bao, L., Wang, L., Cao, L., Chi, H., Hu, Y., Li, Q., Zhou, Y., Jiang, Y., Zhu, Q., Deng, Y., Liu, P., Wang, N., Wang, L., Liu, M., Li, Y., Zhu, B., Fan, K., Fu, W., Yang, P., Pei, X., Cui, Z., Qin, L., Ge, P., Wu, J., Liu, S., Chen, Y., Huang, W., Wang, Q., Qin, C.F., Wang, Y., Qin, C., Wang, X.(2022) Nature 603: 919-925
- PubMed: 35090164 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-04466-x
- Primary Citation Related Structures:
7WE7, 7WE8, 7WE9, 7WEA, 7WEB, 7WEC, 7WED, 7WEE, 7WEF, 7WLC - PubMed Abstract:
Omicron (B.1.1.529), the most heavily mutated SARS-CoV-2 variant so far, is highly resistant to neutralizing antibodies, raising concerns about the effectiveness of antibody therapies and vaccines 1,2 . Here we examined whether sera from individuals who received two or three doses of inactivated SARS-CoV-2 vaccine could neutralize authentic Omicron. The seroconversion rates of neutralizing antibodies were 3.3% (2 out of 60) and 95% (57 out of 60) for individuals who had received 2 and 3 doses of vaccine, respectively. For recipients of three vaccine doses, the geometric mean neutralization antibody titre for Omicron was 16.5-fold lower than for the ancestral virus (254). We isolated 323 human monoclonal antibodies derived from memory B cells in triple vaccinees, half of which recognized the receptor-binding domain, and showed that a subset (24 out of 163) potently neutralized all SARS-CoV-2 variants of concern, including Omicron. Therapeutic treatments with representative broadly neutralizing monoclonal antibodies were highly protective against infection of mice with SARS-CoV-2 Beta (B.1.351) and Omicron. Atomic structures of the Omicron spike protein in complex with three classes of antibodies that were active against all five variants of concern defined the binding and neutralizing determinants and revealed a key antibody escape site, G446S, that confers greater resistance to a class of antibodies that bind on the right shoulder of the receptor-binding domain by altering local conformation at the binding interface. Our results rationalize the use of three-dose immunization regimens and suggest that the fundamental epitopes revealed by these broadly ultrapotent antibodies are rational targets for a universal sarbecovirus vaccine.
- CAS Key Laboratory of Infection and Immunity, National Laboratory of Macromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: