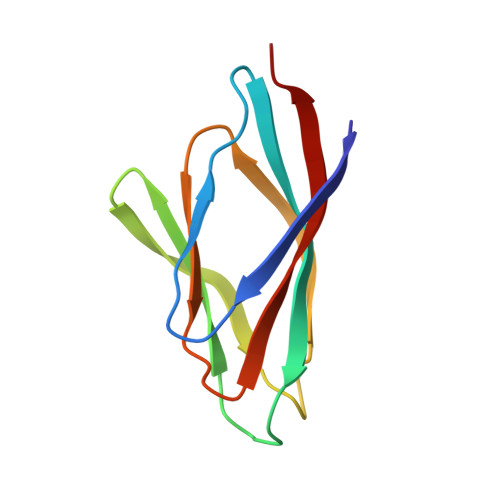
Plant-specific HDT family histone deacetylases are nucleoplasmins.
Bobde, R.C., Kumar, A., Vasudevan, D.(2022) Plant Cell 34: 4760-4777
- PubMed: 36069647 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/plcell/koac275
- Primary Citation Related Structures:
7VMF, 7VMH, 7VMI, 7VRR - PubMed Abstract:
Histone acetyltransferase (HAT)- and histone deacetylase (HDAC)-mediated histone acetylation and deacetylation regulate nucleosome dynamics and gene expression. HDACs are classified into different families, with HD-tuins or HDTs being specific to plants. HDTs show some sequence similarity to nucleoplasmins, the histone chaperones that aid in binding, storing, and loading H2A/H2B dimers to assemble nucleosomes. Here, we solved the crystal structure of the N-terminal domain (NTD) of all four HDTs (HDT1, HDT2, HDT3, and HDT4) from Arabidopsis (Arabidopsis thaliana). The NTDs form a nucleoplasmin fold, exist as pentamers in solution, and are resistant to protease treatment, high temperature, salt, and urea conditions. Structurally, HDTs do not form a decamer, unlike certain classical nucleoplasmins. The HDT-NTD requires an additional A2 acidic tract C-terminal to the nucleoplasmin domain for interaction with histone H3/H4 and H2A/H2B oligomers. We also report the in-solution structures of HDT2 pentamers in complex with histone oligomers. Our study provides a detailed structural and in vitro functional characterization of HDTs, revealing them to be nucleoplasmin family histone chaperones. The experimental confirmation that HDTs are nucleoplasmins may spark new interest in this enigmatic family of proteins.
- Institute of Life Sciences, Bhubaneswar, Odisha 751023, India.
Organizational Affiliation: