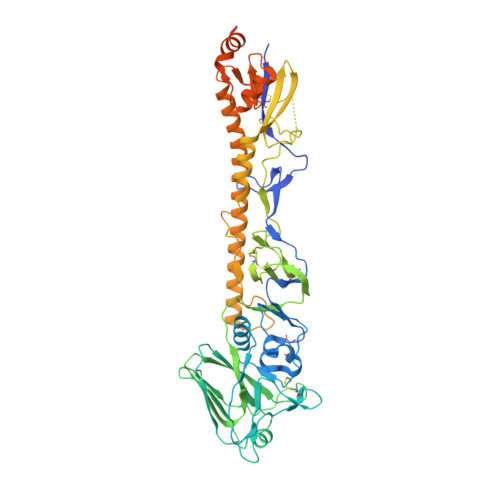
A cryo-electron microscopy support film formed by 2D crystals of hydrophobin HFBI.
Fan, H., Wang, B., Zhang, Y., Zhu, Y., Song, B., Xu, H., Zhai, Y., Qiao, M., Sun, F.(2021) Nat Commun 12: 7257-7257
- PubMed: 34907237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27596-8
- Primary Citation Related Structures:
7VD8, 7VD9, 7VDA, 7VDC, 7VDE, 7VDF - PubMed Abstract:
Cryo-electron microscopy (cryo-EM) has become a powerful tool to resolve high-resolution structures of biomacromolecules in solution. However, air-water interface induced preferred orientations, dissociation or denaturation of biomacromolecules during cryo-vitrification remains a limiting factor for many specimens. To solve this bottleneck, we developed a cryo-EM support film using 2D crystals of hydrophobin HFBI. The hydrophilic side of the HFBI film adsorbs protein particles via electrostatic interactions and sequesters them from the air-water interface, allowing the formation of sufficiently thin ice for high-quality data collection. The particle orientation distribution can be regulated by adjusting the buffer pH. Using this support, we determined the cryo-EM structures of catalase (2.29 Å) and influenza haemagglutinin trimer (2.56 Å), which exhibited strong preferred orientations using a conventional cryo-vitrification protocol. We further show that the HFBI film is suitable to obtain high-resolution structures of small proteins, including aldolase (150 kDa, 3.28 Å) and haemoglobin (64 kDa, 3.6 Å). Our work suggests that HFBI films may have broad future applications in increasing the success rate and efficiency of cryo-EM.
- National Key Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 100101, Beijing, China.
Organizational Affiliation: