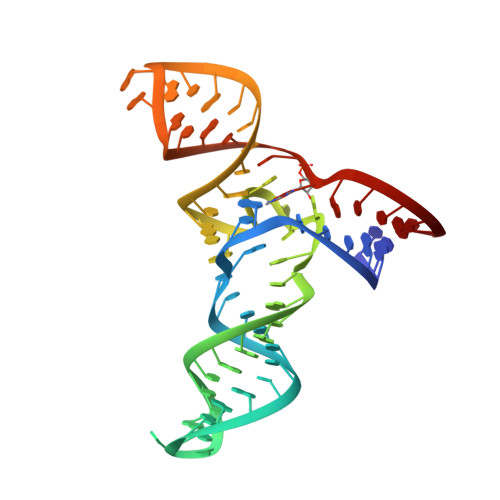
Structure and mechanism of a methyltransferase ribozyme.
Deng, J., Wilson, T.J., Wang, J., Peng, X., Li, M., Lin, X., Liao, W., Lilley, D.M.J., Huang, L.(2022) Nat Chem Biol 18: 556-564
- PubMed: 35301479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-022-00982-z
- Primary Citation Related Structures:
7V9E - PubMed Abstract:
Known ribozymes in contemporary biology perform a limited range of chemical catalysis, but in vitro selection has generated species that catalyze a broader range of chemistry; yet, there have been few structural and mechanistic studies of selected ribozymes. A ribozyme has recently been selected that can catalyze a site-specific methyl transfer reaction. We have solved the crystal structure of this ribozyme at a resolution of 2.3 Å, showing how the RNA folds to generate a very specific binding site for the methyl donor substrate. The structure immediately suggests a catalytic mechanism involving a combination of proximity and orientation and nucleobase-mediated general acid catalysis. The mechanism is supported by the pH dependence of the rate of catalysis. A selected methyltransferase ribozyme can thus use a relatively sophisticated catalytic mechanism, broadening the range of known RNA-catalyzed chemistry.
- Guangdong Provincial Key Laboratory of Malignant Tumor Epigenetics and Gene Regulation, Guangdong-Hong Kong Joint Laboratory for RNA Medicine, Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University, Guangzhou, China.
Organizational Affiliation: