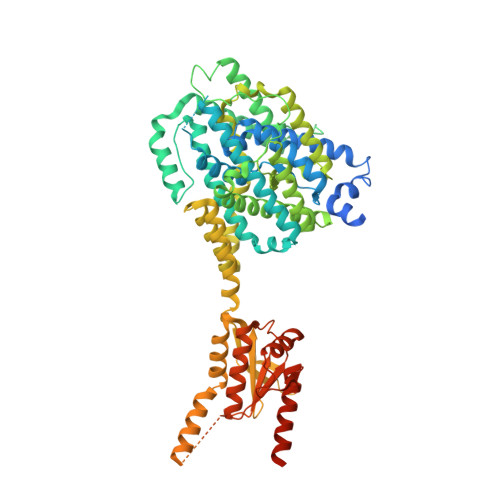
Cryo-EM structures of thermostabilized prestin provide mechanistic insights underlying outer hair cell electromotility.
Futamata, H., Fukuda, M., Umeda, R., Yamashita, K., Tomita, A., Takahashi, S., Shikakura, T., Hayashi, S., Kusakizako, T., Nishizawa, T., Homma, K., Nureki, O.(2022) Nat Commun 13: 6208-6208
- PubMed: 36266333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34017-x
- Primary Citation Related Structures:
7V73, 7V74, 7V75 - PubMed Abstract:
Outer hair cell elecromotility, driven by prestin, is essential for mammalian cochlear amplification. Here, we report the cryo-EM structures of thermostabilized prestin (Pres TS ), complexed with chloride, sulfate, or salicylate at 3.52-3.63 Å resolutions. The central positively-charged cavity allows flexible binding of various anion species, which likely accounts for the known distinct modulations of nonlinear capacitance (NLC) by different anions. Comparisons of these Pres TS structures with recent prestin structures suggest rigid-body movement between the core and gate domains, and provide mechanistic insights into prestin inhibition by salicylate. Mutations at the dimeric interface severely diminished NLC, suggesting that stabilization of the gate domain facilitates core domain movement, thereby contributing to the expression of NLC. These findings advance our understanding of the molecular mechanism underlying mammalian cochlear amplification.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Bunkyo-ku, Tokyo, 113-0033, Japan.
Organizational Affiliation: