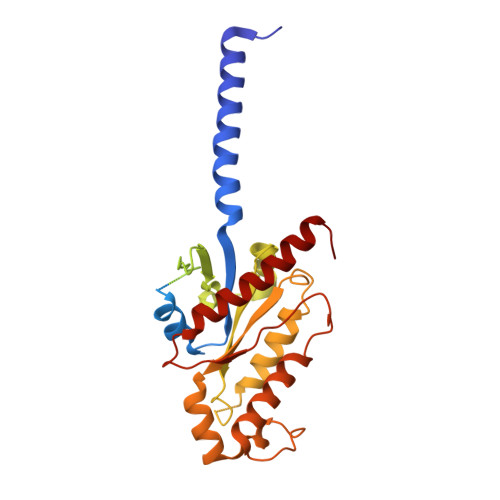
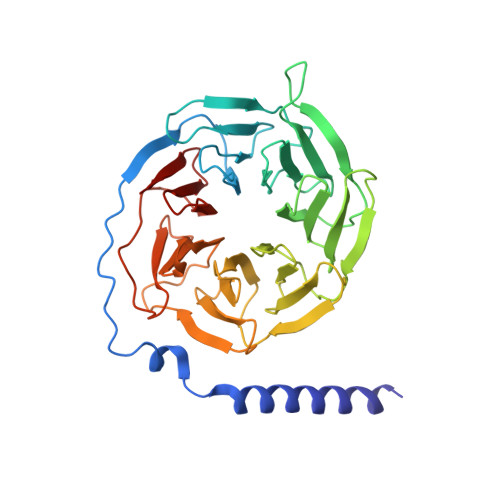
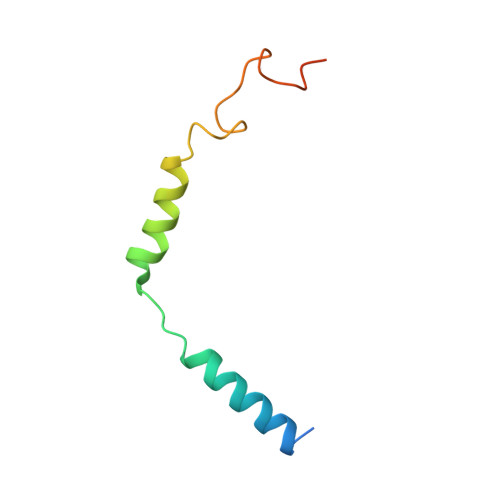
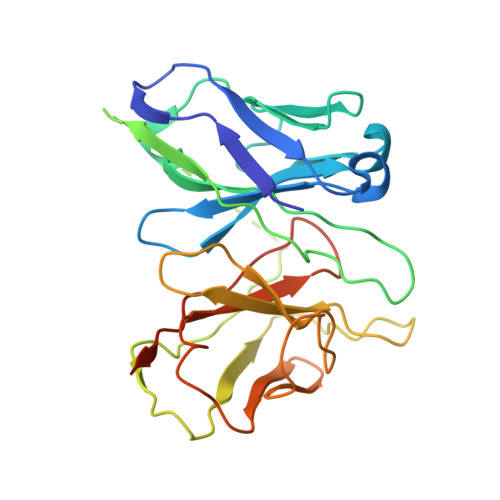
The unconventional activation of the muscarinic acetylcholine receptor M4R by diverse ligands.
Wang, J., Wu, M., Chen, Z., Wu, L., Wang, T., Cao, D., Wang, H., Liu, S., Xu, Y., Li, F., Liu, J., Chen, N., Zhao, S., Cheng, J., Wang, S., Hua, T.(2022) Nat Commun 13: 2855-2855
- PubMed: 35606397 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30595-y
- Primary Citation Related Structures:
7V68, 7V69, 7V6A - PubMed Abstract:
Muscarinic acetylcholine receptors (mAChRs) respond to the neurotransmitter acetylcholine and play important roles in human nervous system. Muscarinic receptor 4 (M4R) is a promising drug target for treating neurological and mental disorders, such as Alzheimer's disease and schizophrenia. However, the lack of understanding on M4R's activation by subtype selective agonists hinders its therapeutic applications. Here, we report the structural characterization of M4R selective allosteric agonist, compound-110, as well as agonist iperoxo and positive allosteric modulator LY2119620. Our cryo-electron microscopy structures of compound-110, iperoxo or iperoxo-LY2119620 bound M4R-G i complex reveal their different interaction modes and activation mechanisms of M4R, and the M4R-ip-LY-G i structure validates the cooperativity between iperoxo and LY2119620 on M4R. Through the comparative structural and pharmacological analysis, compound-110 mostly occupies the allosteric binding pocket with vertical binding pose. Such a binding and activation mode facilitates its allostersic selectivity and agonist profile. In addition, in our schizophrenia-mimic mouse model study, compound-110 shows antipsychotic activity with low extrapyramidal side effects. Thus, this study provides structural insights to develop next-generation antipsychotic drugs selectively targeting on mAChRs subtypes.
- iHuman Institute, ShanghaiTech University, 201210, Shanghai, China.
Organizational Affiliation: