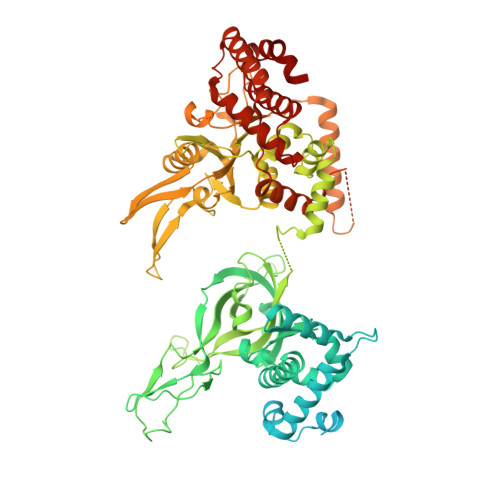
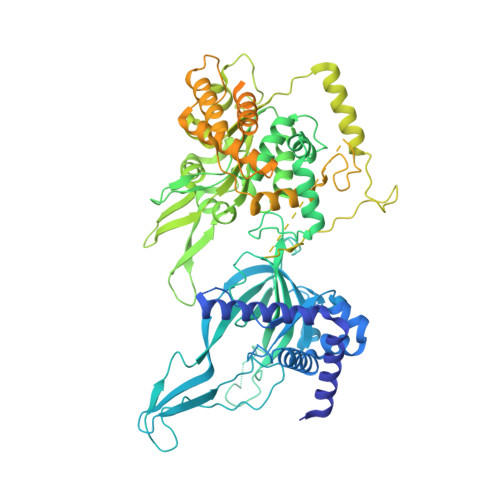
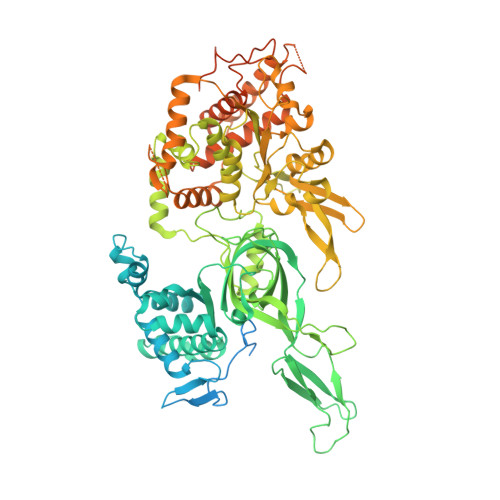
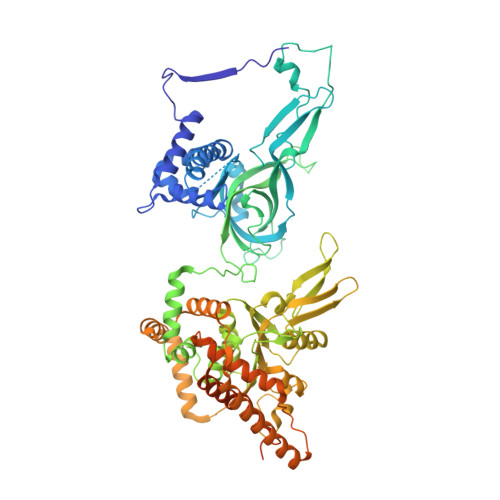
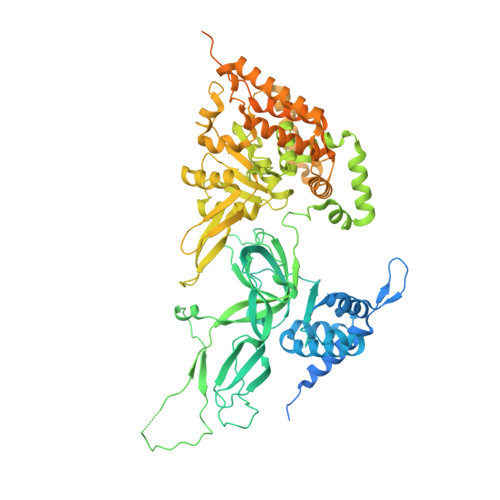
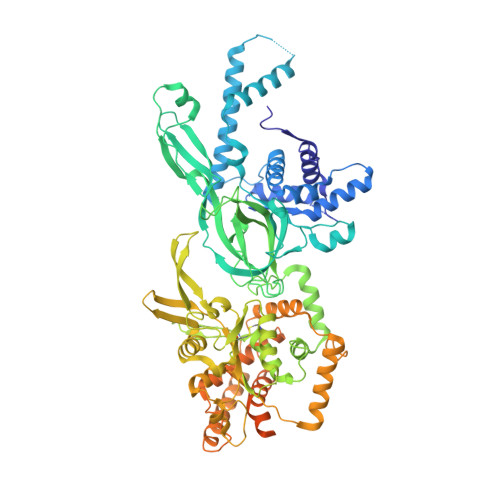
Structural Insight into the MCM double hexamer activation by Dbf4-Cdc7 kinase.
Cheng, J., Li, N., Huo, Y., Dang, S., Tye, B.K., Gao, N., Zhai, Y.(2022) Nat Commun 13: 1396-1396
- PubMed: 35296675 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-29070-5
- Primary Citation Related Structures:
7V3U, 7V3V, 7W8G - PubMed Abstract:
The Dbf4-dependent kinase Cdc7 (DDK) regulates DNA replication initiation by phosphorylation of the MCM double hexamer (MCM-DH) to promote helicase activation. Here, we determine a series of cryo electron microscopy (cryo-EM) structures of yeast DDK bound to the MCM-DH. These structures, occupied by one or two DDKs, differ primarily in the conformations of the kinase core. The interactions of DDK with the MCM-DH are mediated exclusively by subunit Dbf4 straddling across the hexamer interface on the three N-terminal domains (NTDs) of subunits Mcm2, Mcm6, and Mcm4. This arrangement brings Cdc7 close to its only essential substrate, the N-terminal serine/threonine-rich domain (NSD) of Mcm4. Dbf4 further displaces the NSD from its binding site on Mcm4-NTD, facilitating an immediate targeting of this motif by Cdc7. Moreover, the active center of Cdc7 is occupied by a unique Dbf4 inhibitory loop, which is disengaged when the kinase core assumes wobbling conformations. This study elucidates the versatility of Dbf4 in regulating the ordered multisite phosphorylation of the MCM-DH by Cdc7 kinase during helicase activation.
- State Key Laboratory of Membrane Biology, Peking-Tsinghua Joint Center for Life Sciences, School of Life Sciences, Peking University, Beijing, 100871, China.
Organizational Affiliation: