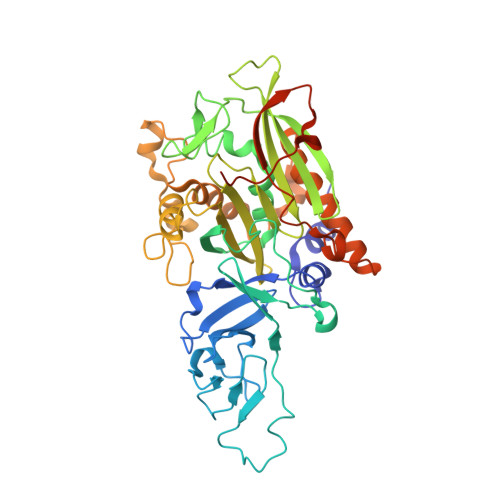
Molecular insights into substrate recognition and catalysis by phthalate dioxygenase from Comamonas testosteroni.
Mahto, J.K., Neetu, N., Waghmode, B., Kuatsjah, E., Sharma, M., Sircar, D., Sharma, A.K., Tomar, S., Eltis, L.D., Kumar, P.(2021) J Biological Chem 297: 101416-101416
- PubMed: 34800435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.101416
- Primary Citation Related Structures:
7FHR, 7FJL, 7V25, 7V28 - PubMed Abstract:
Phthalate, a plasticizer, endocrine disruptor, and potential carcinogen, is degraded by a variety of bacteria. This degradation is initiated by phthalate dioxygenase (PDO), a Rieske oxygenase (RO) that catalyzes the dihydroxylation of phthalate to a dihydrodiol. PDO has long served as a model for understanding ROs despite a lack of structural data. Here we purified PDO KF1 from Comamonas testosteroni KF1 and found that it had an apparent k cat /K m for phthalate of 0.58 ± 0.09 μM -1 s -1 , over 25-fold greater than for terephthalate. The crystal structure of the enzyme at 2.1 Å resolution revealed that it is a hexamer comprising two stacked α 3 trimers, a configuration not previously observed in RO crystal structures. We show that within each trimer, the protomers adopt a head-to-tail configuration typical of ROs. The stacking of the trimers is stabilized by two extended helices, which make the catalytic domain of PDO KF1 larger than that of other characterized ROs. Complexes of PDO KF1 with phthalate and terephthalate revealed that Arg207 and Arg244, two residues on one face of the active site, position these substrates for regiospecific hydroxylation. Consistent with their roles as determinants of substrate specificity, substitution of either residue with alanine yielded variants that did not detectably turnover phthalate. Together, these results provide critical insights into a pollutant-degrading enzyme that has served as a paradigm for ROs and facilitate the engineering of this enzyme for bioremediation and biocatalytic applications.
- Department of Biosciences and Bioengineering, IIT Roorkee, Roorkee, India.
Organizational Affiliation: