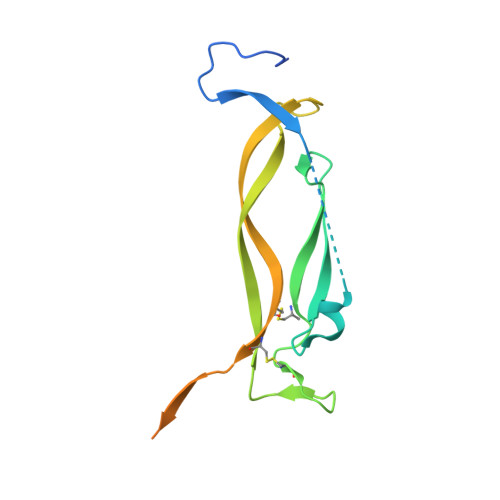
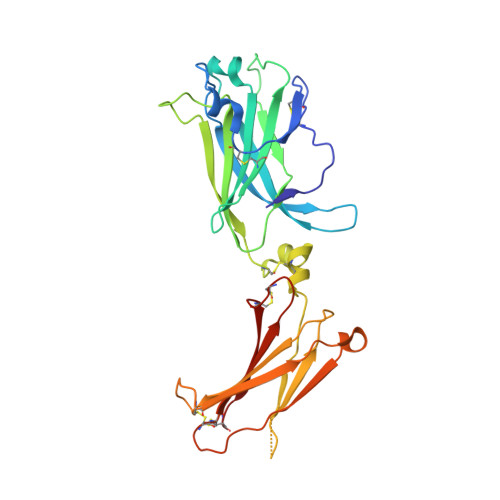
Organizing structural principles of the IL-17 ligand-receptor axis.
Wilson, S.C., Caveney, N.A., Yen, M., Pollmann, C., Xiang, X., Jude, K.M., Hafer, M., Tsutsumi, N., Piehler, J., Garcia, K.C.(2022) Nature 609: 622-629
- PubMed: 35863378 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05116-y
- Primary Citation Related Structures:
7UWJ, 7UWK, 7UWL, 7UWM, 7UWN - PubMed Abstract:
The IL-17 family of cytokines and receptors have central roles in host defence against infection and development of inflammatory diseases 1 . The compositions and structures of functional IL-17 family ligand-receptor signalling assemblies remain unclear. IL-17E (also known as IL-25) is a key regulator of type 2 immune responses and driver of inflammatory diseases, such as allergic asthma, and requires both IL-17 receptor A (IL-17RA) and IL-17RB to elicit functional responses 2 . Here we studied IL-25-IL-17RB binary and IL-25-IL-17RB-IL-17RA ternary complexes using a combination of cryo-electron microscopy, single-molecule imaging and cell-based signalling approaches. The IL-25-IL-17RB-IL-17RA ternary signalling assembly is a C2-symmetric complex in which the IL-25-IL-17RB homodimer is flanked by two 'wing-like' IL-17RA co-receptors through a 'tip-to-tip' geometry that is the key receptor-receptor interaction required for initiation of signal transduction. IL-25 interacts solely with IL-17RB to allosterically promote the formation of the IL-17RB-IL-17RA tip-to-tip interface. The resulting large separation between the receptors at the membrane-proximal level may reflect proximity constraints imposed by the intracellular domains for signalling. Cryo-electron microscopy structures of IL-17A-IL-17RA and IL-17A-IL-17RA-IL-17RC complexes reveal that this tip-to-tip architecture is a key organizing principle of the IL-17 receptor family. Furthermore, these studies reveal dual actions for IL-17RA sharing among IL-17 cytokine complexes, by either directly engaging IL-17 cytokines or alternatively functioning as a co-receptor.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA, USA.
Organizational Affiliation: