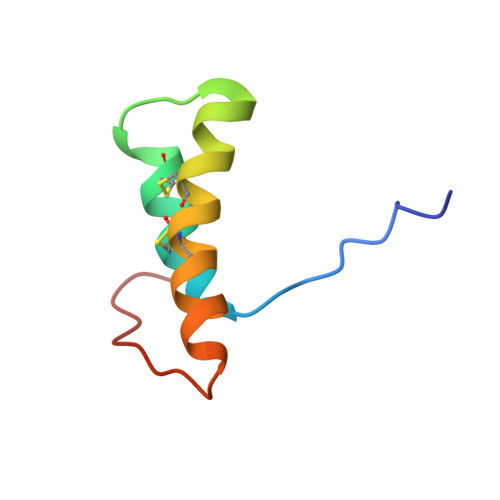
Structure and IgE Cross-Reactivity among Cashew, Pistachio, Walnut, and Peanut Vicilin-Buried Peptides.
Foo, A.C.Y., Nesbit, J.B., Gipson, S.A.Y., DeRose, E.F., Cheng, H., Hurlburt, B.K., Kulis, M.D., Kim, E.H., Dreskin, S.C., Mustafa, S., Maleki, S.J., Mueller, G.A.(2023) J Agric Food Chem 71: 2990-2998
- PubMed: 36728846 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jafc.2c07061
- Primary Citation Related Structures:
7UV1, 7UV2, 7UV3, 7UV4 - PubMed Abstract:
Peanut and tree-nut allergies are frequently comorbid for reasons not completely understood. Vicilin-buried peptides (VBPs) are an emerging family of food allergens whose conserved structural fold could mediate peanut/tree-nut co-allergy. Peptide microarrays were used to identify immunoglobulin E (IgE) epitopes from the N-terminus of the vicilin allergens Ara h 1, Ana o 1, Jug r 2, and Pis v 3 using serum from three patient diagnosis groups: monoallergic to either peanuts or cashew/pistachio, or dual allergic. IgE binding peptides were highly prevalent in the VBP domains AH1.1, AO1.1, JR2.1, and PV3.1, but not in AO1.2, JR2.2, JR2.3, and PV3.2 nor the unstructured regions. The IgE profiles did not correlate with diagnosis group. The structure of the VBPs from cashew and pistachio was solved using solution-NMR. Comparisons of structural features suggest that the VBP scaffold from peanuts and tree-nuts can support cross-reactivity. This may help understand comorbidity and cross-reactivity despite a distant evolutionary origin.
- Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences, 111 T.W. Alexander Drive, MD-MR01, Durham, North Carolina 27709, United States.
Organizational Affiliation: