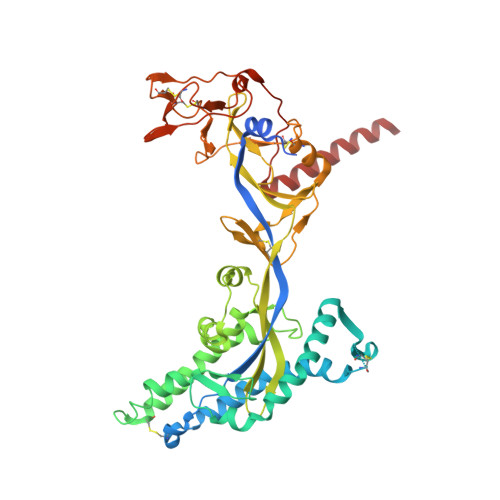
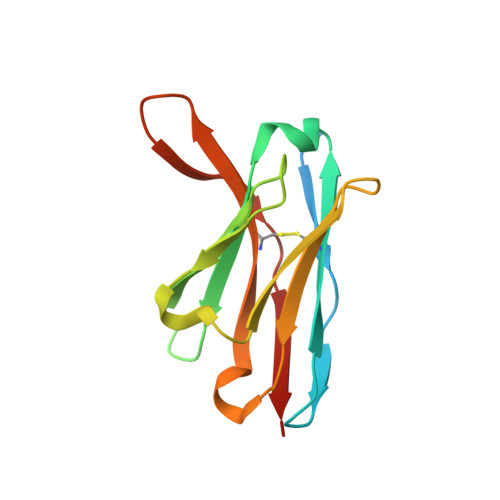
Structural basis for antibody recognition of vulnerable epitopes on Nipah virus F protein.
Byrne, P.O., Fisher, B.E., Ambrozak, D.R., Blade, E.G., Tsybovsky, Y., Graham, B.S., McLellan, J.S., Loomis, R.J.(2023) Nat Commun 14: 1494-1494
- PubMed: 36932063 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36995-y
- Primary Citation Related Structures:
7UOP, 7UP9, 7UPA, 7UPB, 7UPD, 7UPK - PubMed Abstract:
Nipah virus (NiV) is a pathogenic paramyxovirus that causes fatal encephalitis in humans. Two envelope glycoproteins, the attachment protein (G/RBP) and fusion protein (F), facilitate entry into host cells. Due to its vital role, NiV F presents an attractive target for developing vaccines and therapeutics. Several neutralization-sensitive epitopes on the NiV F apex have been described, however the antigenicity of most of the F protein's surface remains uncharacterized. Here, we immunize mice with prefusion-stabilized NiV F and isolate ten monoclonal antibodies that neutralize pseudotyped virus. Cryo-electron microscopy reveals eight neutralization-sensitive epitopes on NiV F, four of which have not previously been described. Novel sites span the lateral and basal faces of NiV F, expanding the known library of vulnerable epitopes. Seven of ten antibodies bind the Hendra virus (HeV) F protein. Multiple sequence alignment suggests that some of these newly identified neutralizing antibodies may also bind F proteins across the Henipavirus genus. This work identifies new epitopes as targets for therapeutics, provides a molecular basis for NiV neutralization, and lays a foundation for development of new cross-reactive antibodies targeting Henipavirus F proteins.
- Department of Molecular Biosciences, The University of Texas at Austin, 78712, Austin, TX, USA.
Organizational Affiliation: