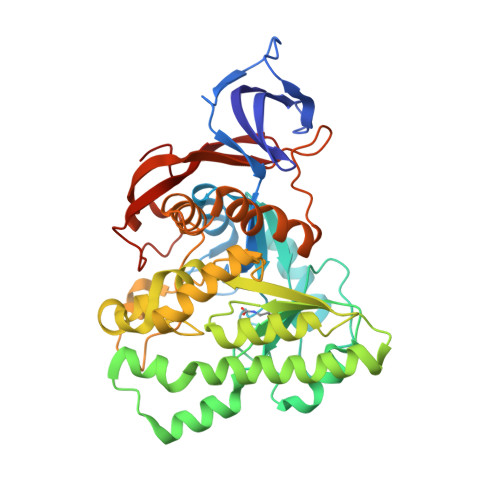
Crystal structure of Methanococcus jannaschii dihydroorotase.
Vitali, J., Nix, J.C., Newman, H.E., Colaneri, M.J.(2023) Proteins 91: 91-98
- PubMed: 35978488 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.26412
- Primary Citation Related Structures:
7UOF - PubMed Abstract:
In this paper, we report the structural analysis of dihydroorotase (DHOase) from the hyperthermophilic and barophilic archaeon Methanococcus jannaschii. DHOase catalyzes the reversible cyclization of N-carbamoyl-l-aspartate to l-dihydroorotate in the third step of de novo pyrimidine biosynthesis. DHOases form a very diverse family of enzymes and have been classified into types and subtypes with structural similarities and differences among them. This is the first archaeal DHOase studied by x-ray diffraction. Its structure and comparison with known representatives of the other subtypes help define the structural features of the archaeal subtype. The M. jannaschii DHOase is found here to have traits from all subtypes. Contrary to expectations, it has a carboxylated lysine bridging the two Zn ions in the active site, and a long catalytic loop. It is a monomeric protein with a large β sandwich domain adjacent to the TIM barrel. Loop 5 is similar to bacterial type III and the C-terminal extension is long.
- Department of Physics, Cleveland State University, Cleveland, Ohio, USA.
Organizational Affiliation: