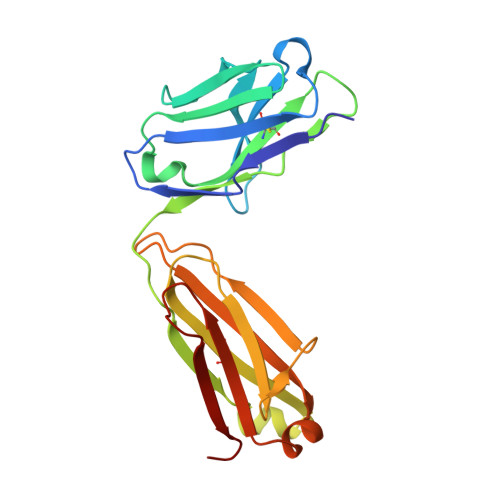
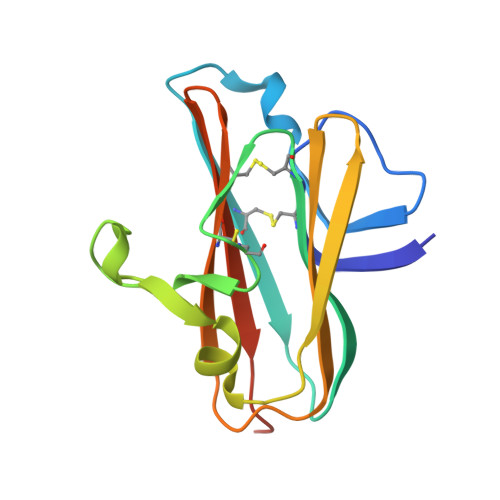
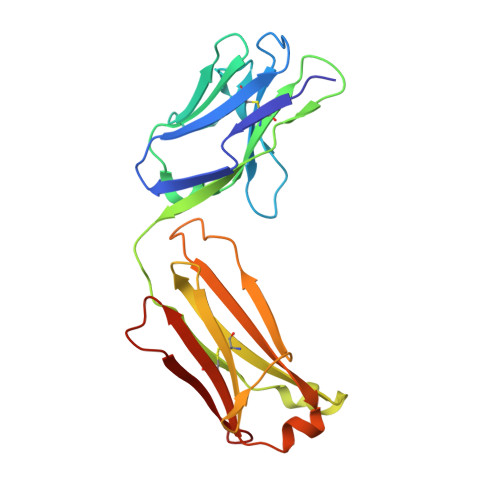
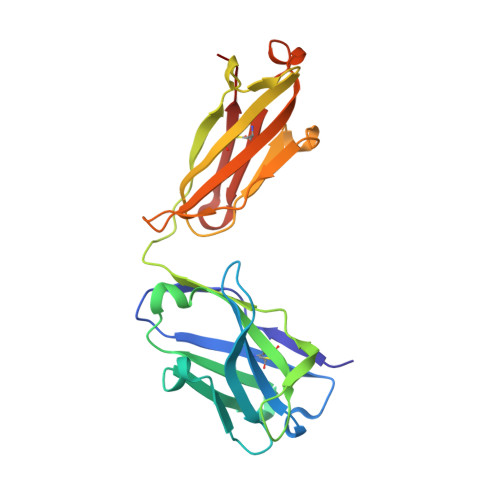
Vaccination with a structure-based stabilized version of malarial antigen Pfs48/45 elicits ultra-potent transmission-blocking antibody responses.
McLeod, B., Mabrouk, M.T., Miura, K., Ravichandran, R., Kephart, S., Hailemariam, S., Pham, T.P., Semesi, A., Kucharska, I., Kundu, P., Huang, W.C., Johnson, M., Blackstone, A., Pettie, D., Murphy, M., Kraft, J.C., Leaf, E.M., Jiao, Y., van de Vegte-Bolmer, M., van Gemert, G.J., Ramjith, J., King, C.R., MacGill, R.S., Wu, Y., Lee, K.K., Jore, M.M., King, N.P., Lovell, J.F., Julien, J.P.(2022) Immunity 55: 1680-1692.e8
- PubMed: 35977542 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2022.07.015
- Primary Citation Related Structures:
7UNB - PubMed Abstract:
Malaria transmission-blocking vaccines (TBVs) aim to elicit human antibodies that inhibit sporogonic development of Plasmodium falciparum in mosquitoes, thereby preventing onward transmission. Pfs48/45 is a leading clinical TBV candidate antigen and is recognized by the most potent transmission-blocking monoclonal antibody (mAb) yet described; still, clinical development of Pfs48/45 antigens has been hindered, largely by its poor biochemical characteristics. Here, we used structure-based computational approaches to design Pfs48/45 antigens stabilized in the conformation recognized by the most potently inhibitory mAb, achieving >25°C higher thermostability compared with the wild-type protein. Antibodies elicited in mice immunized with these engineered antigens displayed on liposome-based or protein nanoparticle-based vaccine platforms exhibited 1-2 orders of magnitude superior transmission-reducing activity, compared with immunogens bearing the wild-type antigen, driven by improved antibody quality. Our data provide the founding principles for using molecular stabilization solely from antibody structure-function information to drive improved immune responses against a parasitic vaccine target.
- Program in Molecular Medicine, The Hospital for Sick Children Research Institute, 686 Bay Street, Toronto, ON M5G 0A4, Canada; Department of Biochemistry, University of Toronto, 1 King's College Circle, Toronto, ON M5S 1A8, Canada.
Organizational Affiliation: