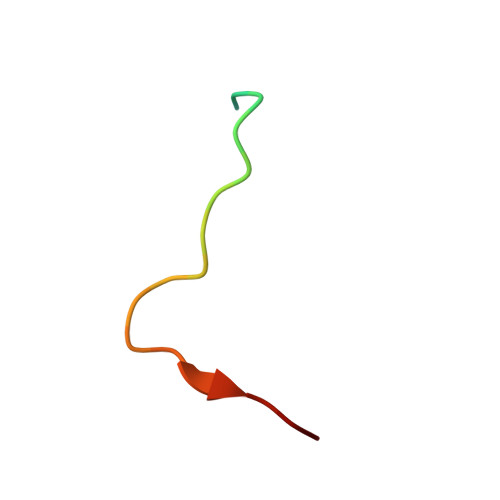
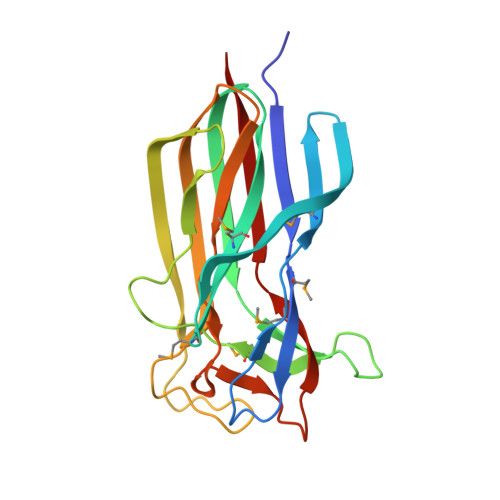
Polerovirus N-terminal readthrough domain structures reveal molecular strategies for mitigating virus transmission by aphids
Schiltz, C.J., Wilson, J.R., Hosford, C.J., Adams, M.C., Preising, S.E., DeBlasio, S.L., MacLeod, H.J., Van Eck, J., Heck, M.L., Chappie, J.S.(2022) Nat Commun 13: 6368