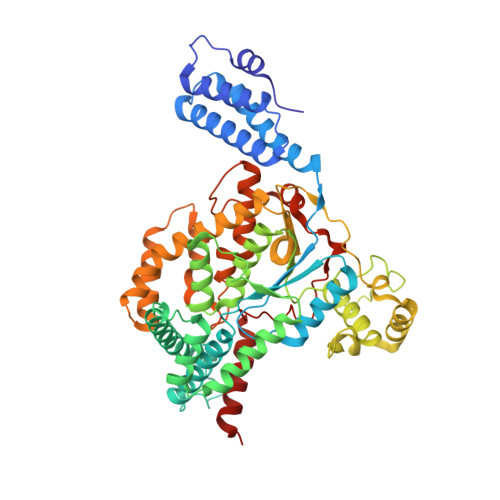
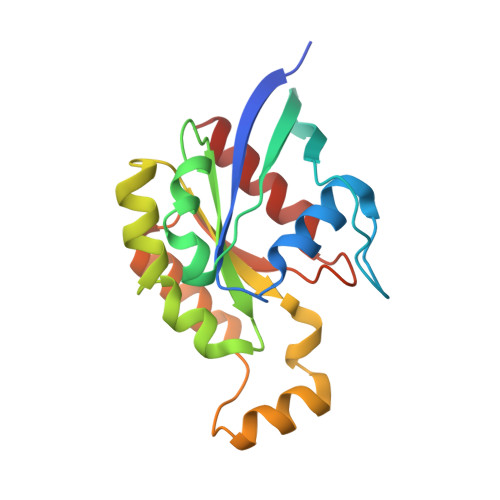
Structure of the glucosyltransferase domain of TcdA in complex with RhoA provides insights into substrate recognition.
Chen, B., Liu, Z., Perry, K., Jin, R.(2022) Sci Rep 12: 9028-9028
- PubMed: 35637242 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-022-12909-8
- Primary Citation Related Structures:
7U2P - PubMed Abstract:
Clostridioides difficile is one of the most common causes of antibiotic-associated diarrhea in developed countries. As key virulence factors of C. difficile, toxin A (TcdA) and toxin B (TcdB) act by glucosylating and inactivating Rho and Ras family small GTPases in host cells, which leads to actin cytoskeleton disruption, cell rounding, and ultimately cell death. Here we present the co-crystal structure of the glucosyltransferase domain (GTD) of TcdA in complex with its substrate human RhoA at 2.60-angstrom resolution. This structure reveals that TcdA GTD grips RhoA mainly through its switch I and switch II regions, which is complemented by interactions involving RhoA's pre-switch I region. Comprehensive structural comparisons between the TcdA GTD-RhoA complex and the structures of TcdB GTD in complex with Cdc42 and R-Ras reveal both the conserved and divergent features of these two toxins in terms of substrate recognition. Taken together, these findings establish the structural basis for TcdA recognition of small GTPases and advance our understanding of the substrates selectivity of large clostridial toxins.
- Department of Physiology and Biophysics, School of Medicine, University of California, Irvine, CA, 92697, USA.
Organizational Affiliation: