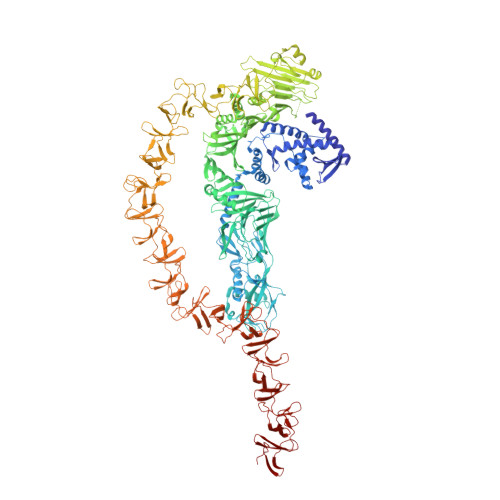Structure and conformational dynamics of Clostridioides difficile toxin A.
Chen, B., Basak, S., Chen, P., Zhang, C., Perry, K., Tian, S., Yu, C., Dong, M., Huang, L., Bowen, M.E., Jin, R.(2022) Life Sci Alliance 5
- PubMed: 35292538 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.202201383
- Primary Citation Related Structures:
7U1Z - PubMed Abstract:
Clostridioides difficile toxin A and B (TcdA and TcdB) are two major virulence factors responsible for diseases associated with C. difficile infection (CDI). Here, we report the 3.18-Å resolution crystal structure of a TcdA fragment (residues L843-T2481), which advances our understanding of the complete structure of TcdA holotoxin. Our structural analysis, together with complementary single molecule FRET and limited proteolysis studies, reveal that TcdA adopts a dynamic structure and its CROPs domain can sample a spectrum of open and closed conformations in a pH-dependent manner. Furthermore, a small globular subdomain (SGS) and the CROPs protect the pore-forming region of TcdA in the closed state at neutral pH, which could contribute to modulating the pH-dependent pore formation of TcdA. A rationally designed TcdA mutation that trapped the CROPs in the closed conformation showed drastically reduced cytotoxicity. Taken together, these studies shed new lights into the conformational dynamics of TcdA and its roles in TcdA intoxication.
- Department of Physiology and Biophysics, School of Medicine, University of California, Irvine, Irvine, CA, USA.
Organizational Affiliation: