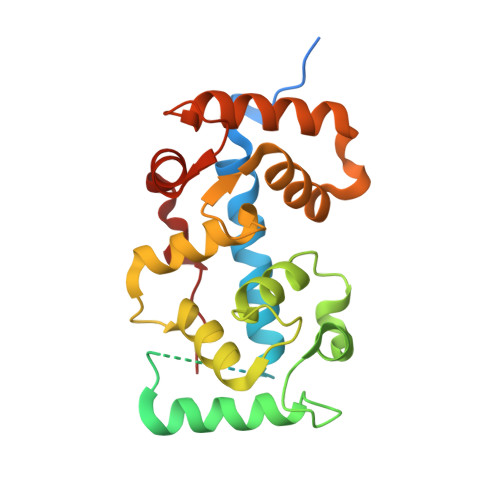
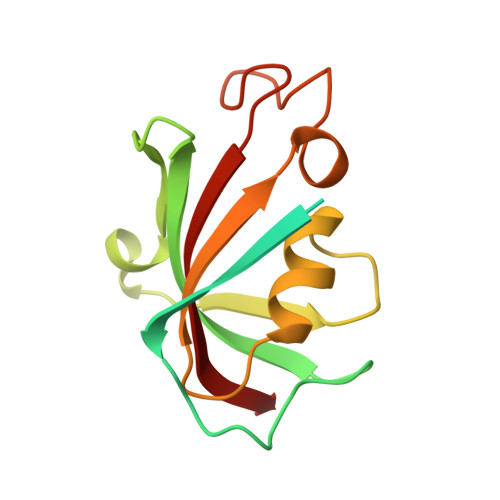
Structure-Guided Synthesis of FK506 and FK520 Analogs with Increased Selectivity Exhibit In Vivo Therapeutic Efficacy against Cryptococcus.
Hoy, M.J., Park, E., Lee, H., Lim, W.Y., Cole, D.C., DeBouver, N.D., Bobay, B.G., Pierce, P.G., Fox 3rd, D., Ciofani, M., Juvvadi, P.R., Steinbach, W., Hong, J., Heitman, J.(2022) mBio 13: e0104922-e0104922
- PubMed: 35604094 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.01049-22
- Primary Citation Related Structures:
7U0S, 7U0T, 7U0U - PubMed Abstract:
Calcineurin is an essential virulence factor that is conserved across human fungal pathogens, including Cryptococcus neoformans, Aspergillus fumigatus, and Candida albicans. Although an excellent target for antifungal drug development, the serine-threonine phosphatase activity of calcineurin is conserved in mammals, and inhibition of this activity results in immunosuppression. FK506 (tacrolimus) is a naturally produced macrocyclic compound that inhibits calcineurin by binding to the immunophilin FKBP12. Previously, our fungal calcineurin-FK506-FKBP12 structure-based approaches identified a nonconserved region of FKBP12 that can be exploited for fungus-specific targeting. These studies led to the design of an FK506 analog, APX879, modified at the C-22 position, which was less immunosuppressive yet maintained antifungal activity. We now report high-resolution protein crystal structures of fungal FKBP12 and a human truncated calcineurin-FKBP12 bound to a natural FK506 analog, FK520 (ascomycin). Based on information from these structures and the success of APX879, we synthesized and screened a novel panel of C-22-modified compounds derived from both FK506 and FK520. One compound, JH-FK-05, demonstrates broad-spectrum antifungal activity in vitro and is nonimmunosuppressive in vivo . In murine models of pulmonary and disseminated C. neoformans infection, JH-FK-05 treatment significantly reduced fungal burden and extended animal survival alone and in combination with fluconazole. Furthermore, molecular dynamic simulations performed with JH-FK-05 binding to fungal and human FKBP12 identified additional residues outside the C-22 and C-21 positions that could be modified to generate novel FK506 analogs with improved antifungal activity. IMPORTANCE Due to rising rates of antifungal drug resistance and a limited armamentarium of antifungal treatments, there is a paramount need for novel antifungal drugs to treat systemic fungal infections. Calcineurin has been established as an essential and conserved virulence factor in several fungi, making it an attractive antifungal target. However, due to the immunosuppressive action of calcineurin inhibitors, they have not been successfully utilized clinically for antifungal treatment in humans. Recent availability of crystal structures of fungal calcineurin-bound inhibitor complexes has enabled the structure-guided design of FK506 analogs and led to a breakthrough in the development of a compound with increased fungal specificity. The development of a calcineurin inhibitor with reduced immunosuppressive activity and maintained therapeutic antifungal activity would add a significant tool to the treatment options for these invasive fungal infections with exceedingly high rates of mortality.
- Department of Molecular Genetics and Microbiology, Duke Universitygrid.26009.3d Medical Center, Durham, North Carolina, USA.
Organizational Affiliation: