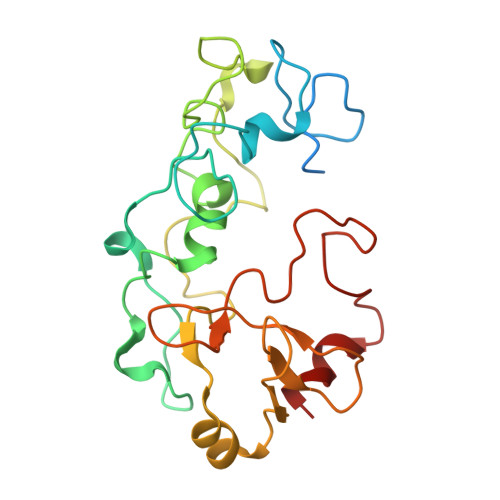
Cryo-EM structure of an extracellular Geobacter OmcE cytochrome filament reveals tetrahaem packing.
Wang, F., Mustafa, K., Suciu, V., Joshi, K., Chan, C.H., Choi, S., Su, Z., Si, D., Hochbaum, A.I., Egelman, E.H., Bond, D.R.(2022) Nat Microbiol 7: 1291-1300
- PubMed: 35798889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-022-01159-z
- Primary Citation Related Structures:
7TFS, 7TGG - PubMed Abstract:
Electrically conductive appendages from the anaerobic bacterium Geobacter sulfurreducens were first observed two decades ago, with genetic and biochemical data suggesting that conductive fibres were type IV pili. Recently, an extracellular conductive filament of G. sulfurreducens was found to contain polymerized c-type cytochrome OmcS subunits, not pilin subunits. Here we report that G. sulfurreducens also produces a second, thinner appendage comprised of cytochrome OmcE subunits and solve its structure using cryo-electron microscopy at ~4.3 Å resolution. Although OmcE and OmcS subunits have no overall sequence or structural similarities, upon polymerization both form filaments that share a conserved haem packing arrangement in which haems are coordinated by histidines in adjacent subunits. Unlike OmcS filaments, OmcE filaments are highly glycosylated. In extracellular fractions from G. sulfurreducens, we detected type IV pili comprising PilA-N and -C chains, along with abundant B-DNA. OmcE is the second cytochrome filament to be characterized using structural and biophysical methods. We propose that there is a broad class of conductive bacterial appendages with conserved haem packing (rather than sequence homology) that enable long-distance electron transport to chemicals or other microbial cells.
- Department of Biochemistry and Molecular Genetics, University of Virginia School of Medicine, Charlottesville, VA, USA.
Organizational Affiliation: