Development and characterization of functional antibodies targeting NMDA receptors.
Tajima, N., Simorowski, N., Yovanno, R.A., Regan, M.C., Michalski, K., Gomez, R., Lau, A.Y., Furukawa, H.(2022) Nat Commun 13: 923-923
- PubMed: 35177668 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28559-3
- Primary Citation Related Structures:
7TE4, 7TE6, 7TE9, 7TEB, 7TEE, 7TEQ, 7TER, 7TES, 7TET - PubMed Abstract:
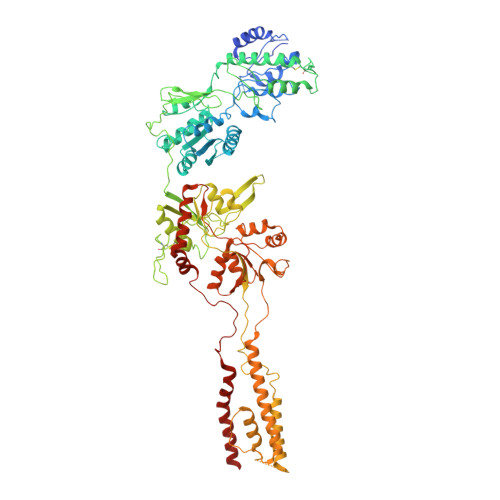
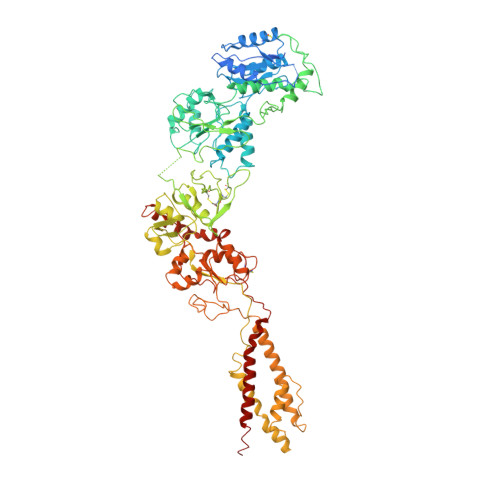
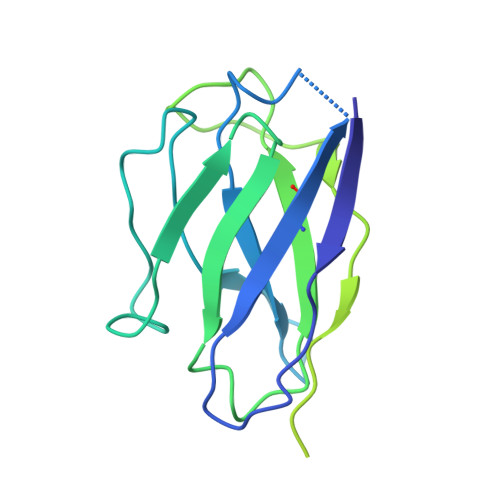
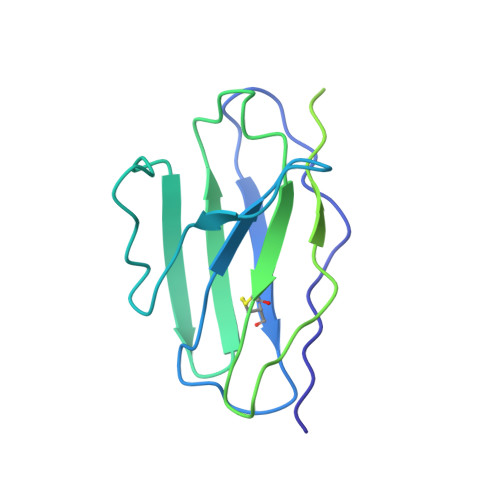
N-methyl-D-aspartate receptors (NMDARs) are critically involved in basic brain functions and neurodegeneration as well as tumor invasiveness. Targeting specific subtypes of NMDARs with distinct activities has been considered an effective therapeutic strategy for neurological disorders and diseases. However, complete elimination of off-target effects of small chemical compounds has been challenging and thus, there is a need to explore alternative strategies for targeting NMDAR subtypes. Here we report identification of a functional antibody that specifically targets the GluN1-GluN2B NMDAR subtype and allosterically down-regulates ion channel activity as assessed by electrophysiology. Through biochemical analysis, x-ray crystallography, single-particle electron cryomicroscopy, and molecular dynamics simulations, we show that this inhibitory antibody recognizes the amino terminal domain of the GluN2B subunit and increases the population of the non-active conformational state. The current study demonstrates that antibodies may serve as specific reagents to regulate NMDAR functions for basic research and therapeutic objectives.
- W.M. Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, NY, 11724, USA.
Organizational Affiliation: