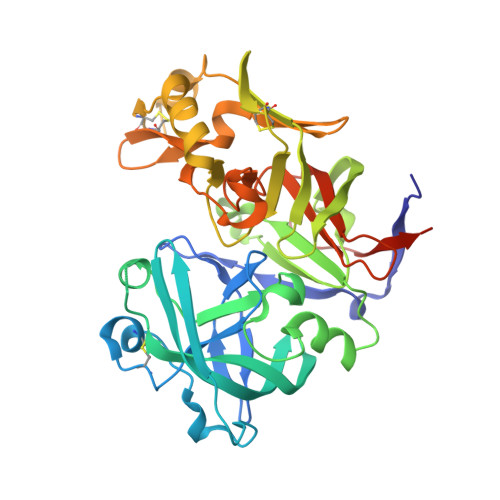
Basis for drug selectivity of plasmepsin IX and X inhibition in Plasmodium falciparum and vivax.
Hodder, A.N., Christensen, J., Scally, S., Triglia, T., Ngo, A., Birkinshaw, R.W., Bailey, B., Favuzza, P., Dietrich, M.H., Tham, W.H., Czabotar, P.E., Lowes, K., Guo, Z., Murgolo, N., Lera Ruiz, M., McCauley, J.A., Sleebs, B.E., Olsen, D., Cowman, A.F.(2022) Structure 30: 947
- PubMed: 35460613 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2022.03.018
- Primary Citation Related Structures:
7TBB, 7TBC, 7TBD, 7TBE - PubMed Abstract:
Plasmepsins IX (PMIX) and X (PMX) are essential aspartyl proteases for Plasmodium spp. egress, invasion, and development. WM4 and WM382 inhibit PMIX and PMX in Plasmodium falciparum and P. vivax. WM4 inhibits PMX, while WM382 is a dual inhibitor of PMIX and PMX. To understand their function, we identified protein substrates. Enzyme kinetic and structural analyses identified interactions responsible for drug specificity. PMIX and PMX have similar substrate specificity; however, there are distinct differences for peptide and protein substrates. Differences in WM4 and WM382 binding for PMIX and PMX map to variations in the S' region and engagement of the active site S3 pocket. Structures of PMX reveal interactions and mechanistic detail of drug binding important for development of clinical candidates against these targets.
- The Walter and Eliza Hall Institute of Medical Research, Parkville 3052 VIC, Australia; University of Melbourne, Melbourne, 3010 VIC, Australia.
Organizational Affiliation: