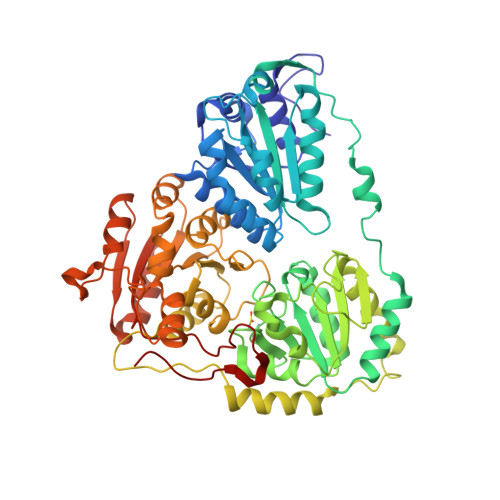
Structural basis of resistance to herbicides that target acetohydroxyacid synthase.
Lonhienne, T., Cheng, Y., Garcia, M.D., Hu, S.H., Low, Y.S., Schenk, G., Williams, C.M., Guddat, L.W.(2022) Nat Commun 13: 3368-3368
- PubMed: 35690625 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-31023-x
- Primary Citation Related Structures:
7STQ, 7TZZ, 7U1D, 7U1U, 7U25 - PubMed Abstract:
Acetohydroxyacid synthase (AHAS) is the target for more than 50 commercial herbicides; first applied to crops in the 1980s. Since then, 197 site-of-action resistance isolates have been identified in weeds, with mutations at P197 and W574 the most prevalent. Consequently, AHAS is at risk of not being a useful target for crop protection. To develop new herbicides, a functional understanding to explain the effect these mutations have on activity is required. Here, we show that these mutations can have two effects (i) to reduce binding affinity of the herbicides and (ii) to abolish time-dependent accumulative inhibition, critical to the exceptional effectiveness of this class of herbicide. In the two mutants, conformational changes occur resulting in a loss of accumulative inhibition by most herbicides. However, bispyribac, a bulky herbicide is able to counteract the detrimental effects of these mutations, explaining why no site-of-action resistance has yet been reported for this herbicide.
- School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, QLD, 4072, Australia.
Organizational Affiliation: