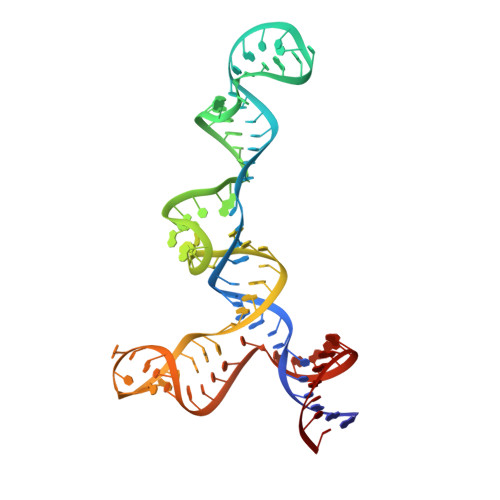
A functional SNP regulates E-cadherin expression by dynamically remodeling the 3D structure of a promoter-associated non-coding RNA transcript.
Sharma, S., Pisignano, G., Merulla, J., Catapano, C.V., Varani, G.(2022) Nucleic Acids Res 50: 11331-11343
- PubMed: 36243981
- DOI: https://doi.org/10.1093/nar/gkac875
- Primary Citation of Related Structures:
7SHX - PubMed Abstract:
Transcription of E-cadherin, a tumor suppressor that plays critical roles in cell adhesion and the epithelial-mesenchymal transition, is regulated by a promoter-associated non-coding RNA (paRNA). The sense-oriented paRNA (S-paRNA) includes a functional C/A single nucleotide polymorphism (SNP rs16260). The A-allele leads to decreased transcriptional activity and increased prostate cancer risk. The polymorphic site is known to affect binding of a microRNA-guided Argonaute 1 (AGO1) complex and recruitment of chromatin-modifying enzymes to silence the promoter. Yet the SNP is distant from the microRNA-AGO1 binding domain in both primary sequence and secondary structure, raising the question of how regulation occurs. Here we report the 3D NMR structure of the 104-nucleotide domain of the S-paRNA that encompasses the SNP and the microRNA-binding site. We show that the A to C change alters the locally dynamic and metastable structure of the S-paRNA, revealing how the single nucleotide mutation regulates the E-cadherin promoter through its effect on the non-coding RNA structure.
Organizational Affiliation:
Department of Chemistry, University of Washington, Seattle, WA 98195-1700, USA.