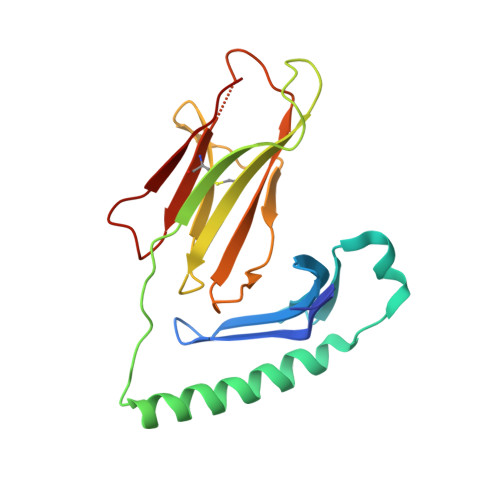
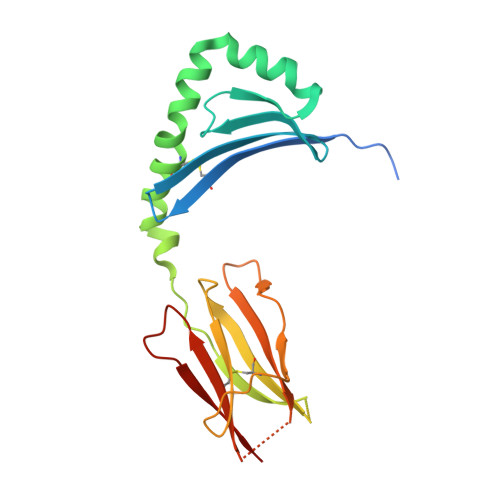
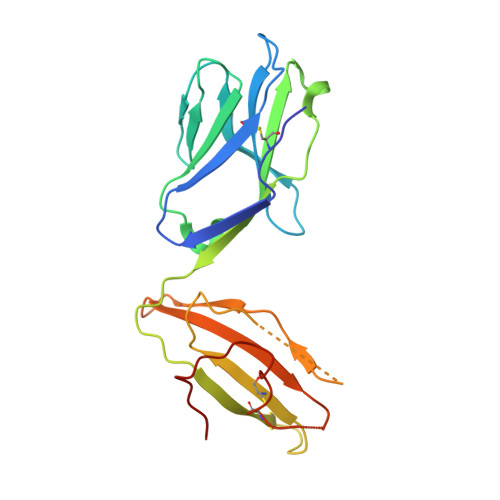
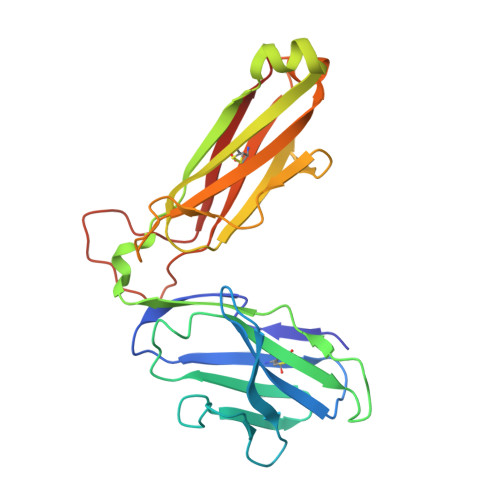
Structural basis of T cell receptor specificity and cross-reactivity of two HLA-DQ2.5-restricted gluten epitopes in celiac disease.
Ciacchi, L., Farenc, C., Dahal-Koirala, S., Petersen, J., Sollid, L.M., Reid, H.H., Rossjohn, J.(2022) J Biological Chem 298: 101619-101619
- PubMed: 35065967 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.101619
- Primary Citation Related Structures:
7SG0, 7SG1, 7SG2 - PubMed Abstract:
Celiac disease is a T cell-mediated chronic inflammatory condition often characterized by human leukocyte antigen (HLA)-DQ2.5 molecules presenting gluten epitopes derived from wheat, barley, and rye. Although some T cells exhibit cross-reactivity toward distinct gluten epitopes, the structural basis underpinning such cross-reactivity is unclear. Here, we investigated the T-cell receptor specificity and cross-reactivity of two immunodominant wheat gluten epitopes, DQ2.5-glia-α1a (PFPQPELPY) and DQ2.5-glia-ω1 (PFPQPEQPF). We show by surface plasmon resonance that a T-cell receptor alpha variable (TRAV) 4 + -T-cell receptor beta variable (TRBV) 29-1 + TCR bound to HLA-DQ2.5-glia-α1a and HLA-DQ2.5-glia-ω1 with similar affinity, whereas a TRAV4 - (TRAV9-2 + ) TCR recognized HLA-DQ2.5-glia-ω1 only. We further determined the crystal structures of the TRAV4 + -TRBV29-1 + TCR bound to HLA-DQ2.5-glia-α1a and HLA-DQ2.5-glia-ω1, as well as the structure of an epitope-specific TRAV9-2 + -TRBV7-3 + TCR-HLA-DQ2.5-glia-ω1 complex. We found that position 7 (p7) of the DQ2.5-glia-α1a and DQ2.5-glia-ω1 epitopes made very limited contacts with the TRAV4 + TCR, thereby explaining the TCR cross-reactivity across these two epitopes. In contrast, within the TRAV9-2 + TCR-HLA-DQ2.5-glia-ω1 ternary complex, the p7-Gln was situated in an electrostatic pocket formed by the hypervariable CDR3β loop of the TCR and Arg70β from HLA-DQ2.5, a polar network which would not be supported by the p7-Leu residue of DQ2.5-glia-α1a. In conclusion, we provide additional insights into the molecular determinants of TCR specificity and cross-reactivity to two closely-related epitopes in celiac disease.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: