A Series of Novel, Highly Potent, and Orally Bioavailable Next-Generation Tricyclic Peptide PCSK9 Inhibitors.
Tucker, T.J., Embrey, M.W., Alleyne, C., Amin, R.P., Bass, A., Bhatt, B., Bianchi, E., Branca, D., Bueters, T., Buist, N., Ha, S.N., Hafey, M., He, H., Higgins, J., Johns, D.G., Kerekes, A.D., Koeplinger, K.A., Kuethe, J.T., Li, N., Murphy, B., Orth, P., Salowe, S., Shahripour, A., Tracy, R., Wang, W., Wu, C., Xiong, Y., Zokian, H.J., Wood, H.B., Walji, A.(2021) J Med Chem 64: 16770-16800
- PubMed: 34704436 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01599
- Primary Citation Related Structures:
7S5G, 7S5H - PubMed Abstract:
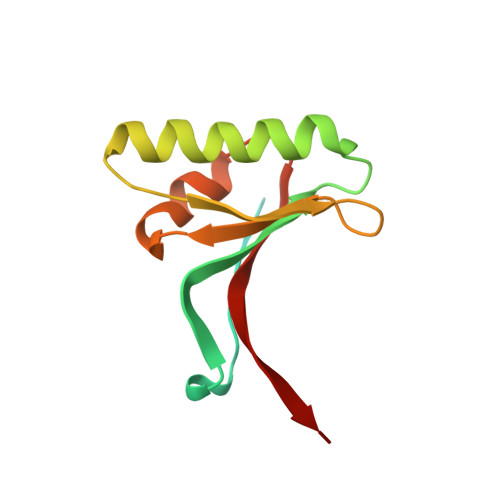
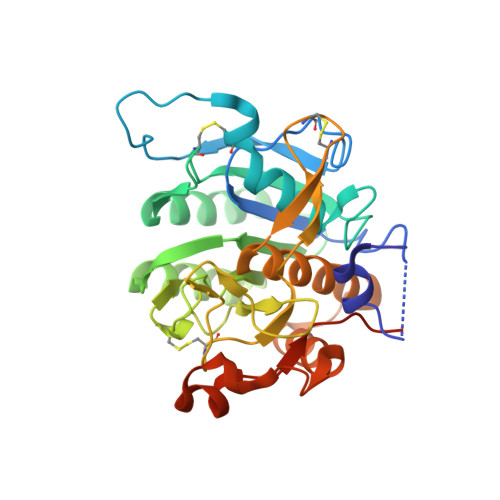
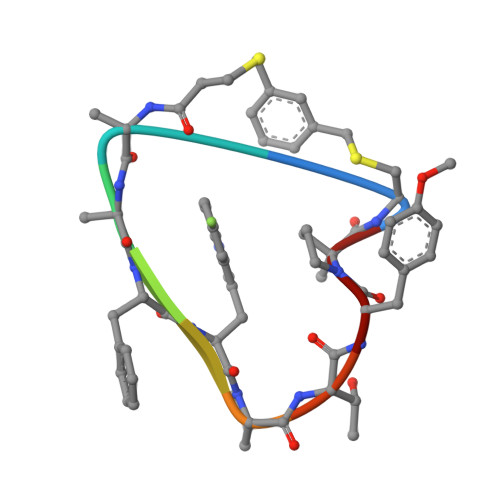
Proprotein convertase subtilisin-like/kexin type 9 (PCSK9) is a key regulator of plasma LDL-cholesterol (LDL-C) and a clinically validated target for the treatment of hypercholesterolemia and coronary artery disease. Starting from second-generation lead structures such as 2 , we were able to refine these structures to obtain extremely potent bi- and tricyclic PCSK9 inhibitor peptides. Optimized molecules such as 44 demonstrated sufficient oral bioavailability to maintain therapeutic levels in rats and cynomolgus monkeys after dosing with an enabled formulation. We demonstrated target engagement and LDL lowering in cynomolgus monkeys essentially identical to those observed with the clinically approved, parenterally dosed antibodies. These molecules represent the first report of highly potent and orally bioavailable macrocyclic peptide PCSK9 inhibitors with overall profiles favorable for potential development as once-daily oral lipid-lowering agents. In this manuscript, we detail the design criteria and multiparameter optimization of this novel series of PCSK9 inhibitors.
- Department of Medicinal Chemistry, Merck & Co., Inc., 770 Sumneytown Pike, P.O. Box 4, West Point, Pennsylvania 19486 United States.
Organizational Affiliation: