Molecular basis for integrin adhesion receptor binding to p21-activated kinase 4 (PAK4)
Ha, B.H., Yigit, S., Natarajan, N., Morse, E.M., Calderwood, D.A., Boggon, T.J.(2022) Commun Biol 5: 1257
Experimental Data Snapshot
Starting Model: experimental
View more details
(2022) Commun Biol 5: 1257
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Isoform 2 of Serine/threonine-protein kinase PAK 4 | 346 | Homo sapiens | Mutation(s): 2 Gene Names: PAK4, KIAA1142 EC: 2.7.11.1 | 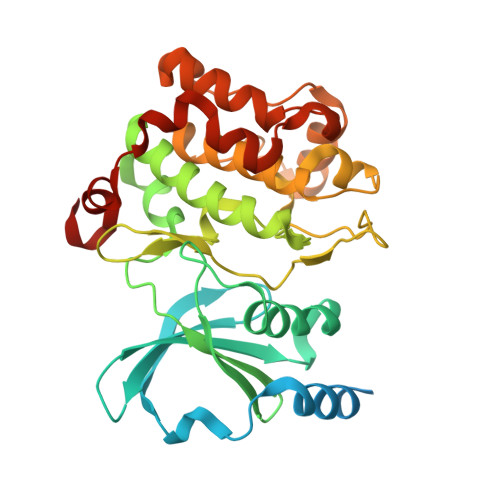 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O96013 GTEx: ENSG00000130669 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O96013 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Integrin beta-5 | 11 | Homo sapiens | Mutation(s): 0 | 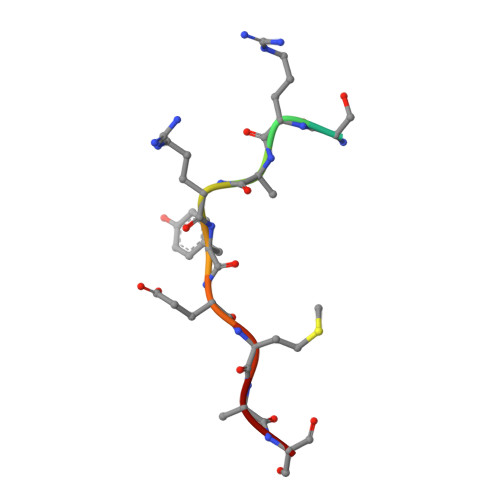 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P18084 GTEx: ENSG00000082781 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P18084 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ANP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A] | PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER C10 H17 N6 O12 P3 PVKSNHVPLWYQGJ-KQYNXXCUSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 61.821 | α = 90 |
| b = 61.821 | β = 90 |
| c = 178.829 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | R01GM138411 |