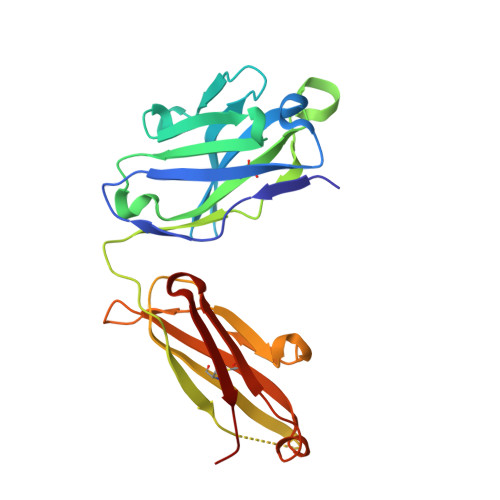
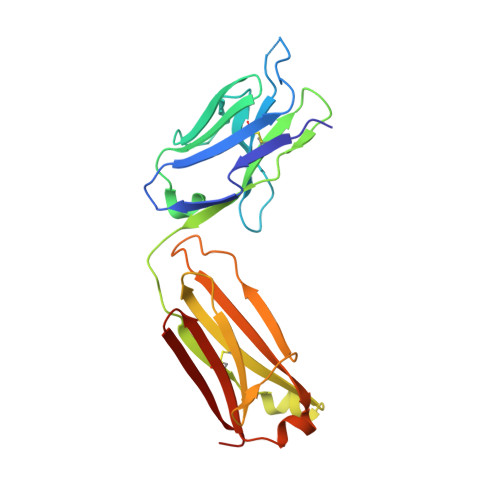
Stabilized coronavirus spike stem elicits a broadly protective antibody.
Hsieh, C.L., Werner, A.P., Leist, S.R., Stevens, L.J., Falconer, E., Goldsmith, J.A., Chou, C.W., Abiona, O.M., West, A., Westendorf, K., Muthuraman, K., Fritch, E.J., Dinnon 3rd, K.H., Schafer, A., Denison, M.R., Chappell, J.D., Baric, R.S., Graham, B.S., Corbett, K.S., McLellan, J.S.(2021) Cell Rep 37: 109929-109929
- PubMed: 34710354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2021.109929
- Primary Citation Related Structures:
7S3M, 7S3N - PubMed Abstract:
Current coronavirus (CoV) vaccines primarily target immunodominant epitopes in the S1 subunit, which are poorly conserved and susceptible to escape mutations, thus threatening vaccine efficacy. Here, we use structure-guided protein engineering to remove the S1 subunit from the Middle East respiratory syndrome (MERS)-CoV spike (S) glycoprotein and develop stabilized stem (SS) antigens. Vaccination with MERS SS elicits cross-reactive β-CoV antibody responses and protects mice against lethal MERS-CoV challenge. High-throughput screening of antibody-secreting cells from MERS SS-immunized mice led to the discovery of a panel of cross-reactive monoclonal antibodies. Among them, antibody IgG22 binds with high affinity to both MERS-CoV and severe acute respiratory syndrome (SARS)-CoV-2 S proteins, and a combination of electron microscopy and crystal structures localizes the epitope to a conserved coiled-coil region in the S2 subunit. Passive transfer of IgG22 protects mice against both MERS-CoV and SARS-CoV-2 challenge. Collectively, these results provide a proof of principle for cross-reactive CoV antibodies and inform the development of pan-CoV vaccines and therapeutic antibodies.
- Department of Molecular Biosciences, University of Texas at Austin, Austin, TX 78712, USA.
Organizational Affiliation: