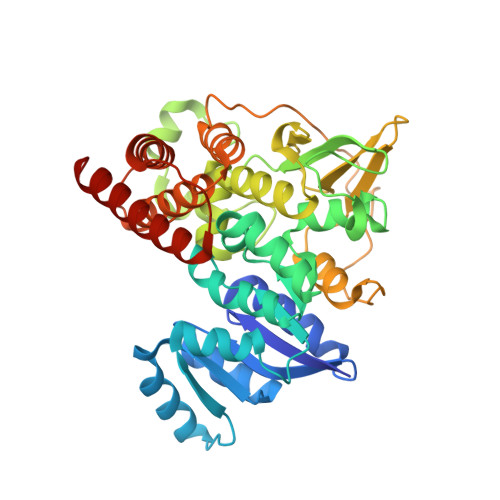
Characterization and Inhibition of 1-Deoxy-d-Xylulose 5-Phosphate Reductoisomerase: A Promising Drug Target in Acinetobacter baumannii and Klebsiella pneumoniae .
Ball, H.S., Girma, M.B., Zainab, M., Soojhawon, I., Couch, R.D., Noble, S.M.(2021) ACS Infect Dis 7: 2987-2998
- PubMed: 34672535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.1c00132
- Primary Citation Related Structures:
7S04 - PubMed Abstract:
The ESKAPE pathogens comprise a group of multidrug-resistant bacteria that are the leading cause of nosocomial infections worldwide. The prevalence of antibiotic resistant strains and the relative ease by which bacteria acquire resistance genes highlight the continual need for the development of novel antibiotics against new drug targets. The methylerythritol phosphate (MEP) pathway is an attractive target for the development of new antibiotics. The MEP pathway governs the synthesis of isoprenoids, which are key lipid precursors for vital cell components such as ubiquinone and bacterial hopanoids. Additionally, the MEP pathway is entirely distinct from the corresponding mammalian pathway, the mevalonic acid (MVA) pathway, making the first committed enzyme of the MEP pathway, 1-deoxy-d-xylulose 5-phosphate reductoisomerase (IspC), an attractive target for antibiotic development. To facilitate drug development against two of the ESKAPE pathogens, Acinetobacter baumannii and Klebsiella pneumoniae , we cloned, expressed, purified, and characterized IspC from these two Gram-negative bacteria. Enzyme inhibition assays using IspC from these two pathogens, and compounds fosmidomycin and FR900098, indicate IC 50 values ranging from 19.5-45.5 nM. Antimicrobial susceptibility tests with these inhibitors reveal that A. baumannii is susceptible to FR900098, whereas K. pneumoniae is susceptible to both compounds. Finally, to facilitate structure-based drug design of inhibitors targeting A. baumannii IspC, we determined the 2.5 Å crystal structure of IspC from A. baumannii in complex with inhibitor FR900098, and cofactors NADPH and magnesium.
- Department of Chemistry and Biochemistry, George Mason University, Manassas, Virginia 20109, United States of America.
Organizational Affiliation: