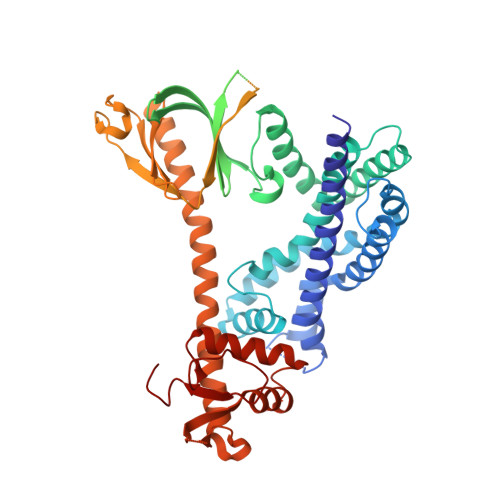
Structure of the metastatic factor P-Rex1 reveals a two-layered autoinhibitory mechanism.
Chang, Y.G., Lupton, C.J., Bayly-Jones, C., Keen, A.C., D'Andrea, L., Lucato, C.M., Steele, J.R., Venugopal, H., Schittenhelm, R.B., Whisstock, J.C., Halls, M.L., Ellisdon, A.M.(2022) Nat Struct Mol Biol 29: 767-773
- PubMed: 35864164 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00804-9
- Primary Citation Related Structures:
7RX9, 7SYF - PubMed Abstract:
P-Rex (PI(3,4,5)P 3 -dependent Rac exchanger) guanine nucleotide exchange factors potently activate Rho GTPases. P-Rex guanine nucleotide exchange factors are autoinhibited, synergistically activated by Gβγ and PI(3,4,5)P 3 binding and dysregulated in cancer. Here, we use X-ray crystallography, cryogenic electron microscopy and crosslinking mass spectrometry to determine the structural basis of human P-Rex1 autoinhibition. P-Rex1 has a bipartite structure of N- and C-terminal modules connected by a C-terminal four-helix bundle that binds the N-terminal Pleckstrin homology (PH) domain. In the N-terminal module, the Dbl homology (DH) domain catalytic surface is occluded by the compact arrangement of the DH-PH-DEP1 domains. Structural analysis reveals a remarkable conformational transition to release autoinhibition, requiring a 126° opening of the DH domain hinge helix. The off-axis position of Gβγ and PI(3,4,5)P 3 binding sites further suggests a counter-rotation of the P-Rex1 halves by 90° facilitates PH domain uncoupling from the four-helix bundle, releasing the autoinhibited DH domain to drive Rho GTPase signaling.
- Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia. tyler.chang@monash.edu.
Organizational Affiliation: