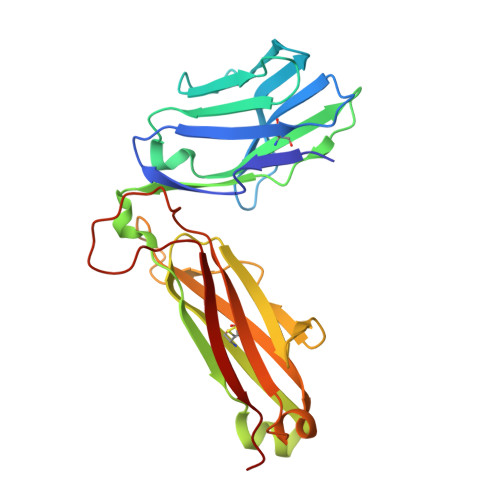
Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR.
Szeto, C., Nguyen, A.T., Lobos, C.A., Chatzileontiadou, D.S.M., Jayasinghe, D., Grant, E.J., Riboldi-Tunnicliffe, A., Smith, C., Gras, S.(2021) Cells 10
- PubMed: 34685626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/cells10102646
- Primary Citation Related Structures:
7RTD, 7RTR - PubMed Abstract:
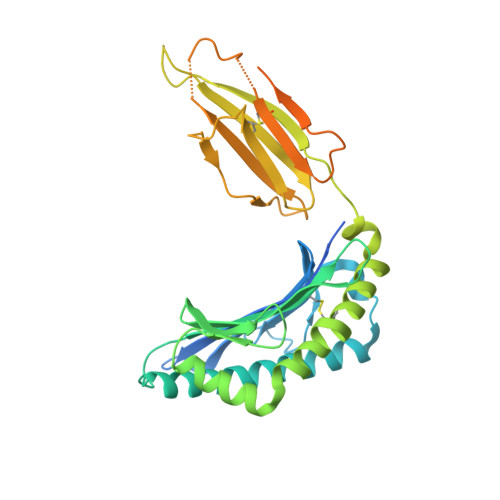
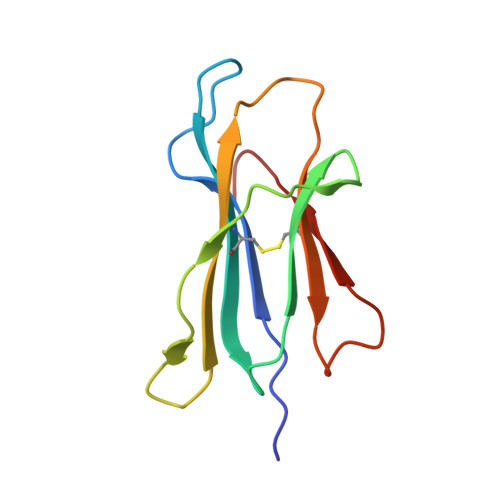
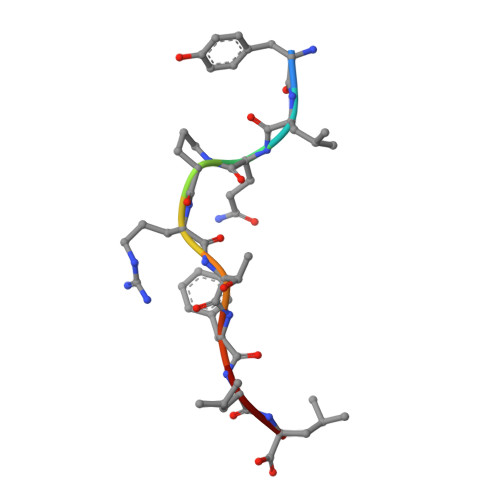
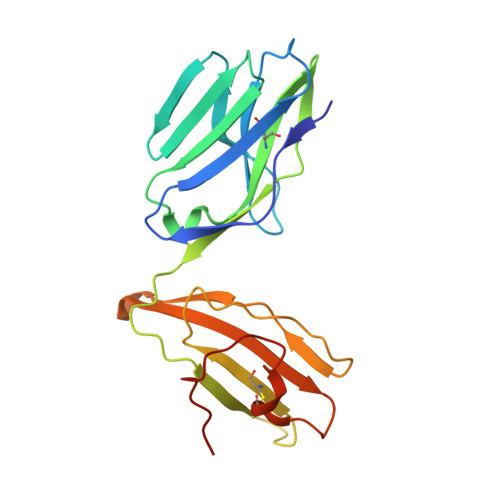
The data currently available on how the immune system recognises the SARS-CoV-2 virus is growing rapidly. While there are structures of some SARS-CoV-2 proteins in complex with antibodies, which helps us understand how the immune system is able to recognise this new virus; however, we lack data on how T cells are able to recognise this virus. T cells, especially the cytotoxic CD8+ T cells, are critical for viral recognition and clearance. Here we report the X-ray crystallography structure of a T cell receptor, shared among unrelated individuals (public TCR) in complex with a dominant spike-derived CD8+ T cell epitope (YLQ peptide). We show that YLQ activates a polyfunctional CD8+ T cell response in COVID-19 recovered patients. We detail the molecular basis for the shared TCR gene usage observed in HLA-A*02:01+ individuals, providing an understanding of TCR recognition towards a SARS-CoV-2 epitope. Interestingly, the YLQ peptide conformation did not change upon TCR binding, facilitating the high-affinity interaction observed.
- Viral and Structural Immunology Laboratory, Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, School of Molecular Sciences, La Trobe University, Bundoora, VIC 3086, Australia.
Organizational Affiliation: