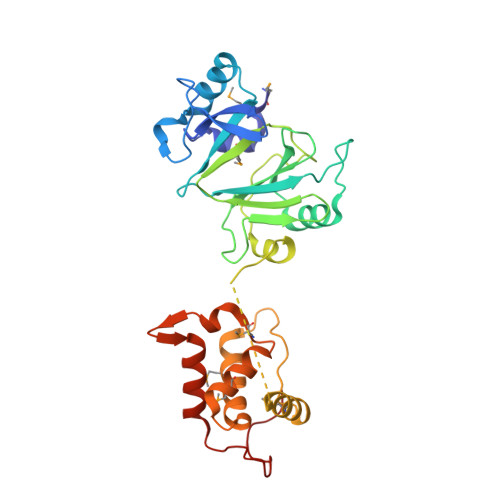
Molecular Dissection of the Primase and Polymerase Activities of Deep-Sea Phage NrS-1 Primase-Polymerase.
Huang, F., Lu, X., Yu, C., Sliz, P., Wang, L., Zhu, B.(2021) Front Microbiol 12: 766612-766612
- PubMed: 34975792 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmicb.2021.766612
- Primary Citation Related Structures:
7RR3 - PubMed Abstract:
PrimPols are a class of primases that belong to the archaeo-eukaryotic primase (AEP) superfamily but have both primase and DNA polymerase activities. Replicative polymerase from NrS-1 phage (NrSPol) is a representative of the PrimPols. In this study, we identified key residues for the catalytic activity of NrSPol and found that a loop in NrSPol functionally replaces the zinc finger motif that is commonly found in other AEP family proteins. A helix bundle domain (HBD), conserved in the AEP superfamily, was recently reported to bind to the primase recognition site and to be crucial for initiation of primer synthesis. We found that NrSPol can recognize different primase recognition sites, and that the initiation site for primer synthesis is not stringent, suggesting that the HBD conformation is flexible. More importantly, we found that although the HBD-inactivating mutation impairs the primase activity of NrSPol, it significantly enhances the DNA polymerase activity, indicating that the HBD hinders the DNA polymerase activity. The conflict between the primase activity and the DNA polymerase activity in a single protein with the same catalytic domain may be one reason for why DNA polymerases are generally unable to synthesize DNA de novo .
- Key Laboratory of Molecular Biophysics, The Ministry of Education, College of Life Science and Technology and Shenzhen College, Huazhong University of Science and Technology, Wuhan, China.
Organizational Affiliation: