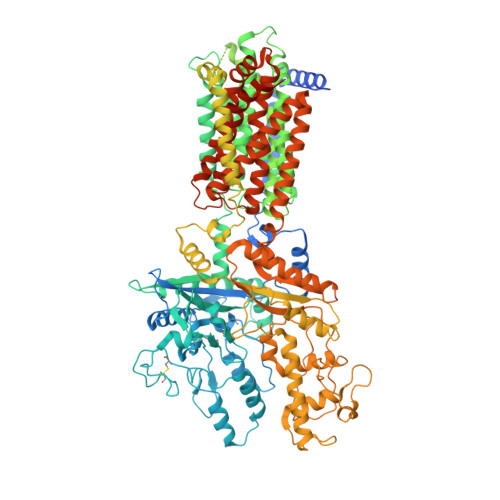
Structural basis for catalyzed assembly of the Sonic hedgehog-Patched1 signaling complex.
Huang, P., Wierbowski, B.M., Lian, T., Chan, C., Garcia-Linares, S., Jiang, J., Salic, A.(2022) Dev Cell 57: 670-685.e8
- PubMed: 35231446 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.devcel.2022.02.008
- Primary Citation Related Structures:
7RHQ, 7RHR - PubMed Abstract:
The dually lipidated Sonic hedgehog (SHH) morphogen signals through the tumor suppressor membrane protein Patched1 (PTCH1) to activate the Hedgehog pathway, which is fundamental in development and cancer. SHH engagement with PTCH1 requires the GAS1 coreceptor, but the mechanism is unknown. We demonstrate a unique role for GAS1, catalyzing SHH-PTCH1 complex assembly in vertebrate cells by direct SHH transfer from the extracellular SCUBE2 carrier to PTCH1. Structure of the GAS1-SHH-PTCH1 transition state identifies how GAS1 recognizes the SHH palmitate and cholesterol modifications in modular fashion and how it facilitates lipid-dependent SHH handoff to PTCH1. Structure-guided experiments elucidate SHH movement from SCUBE2 to PTCH1, explain disease mutations, and demonstrate that SHH-induced PTCH1 dimerization causes its internalization from the cell surface. These results define how the signaling-competent SHH-PTCH1 complex assembles, the key step triggering the Hedgehog pathway, and provide a paradigm for understanding morphogen reception and its regulation.
- Department of Cell Biology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: