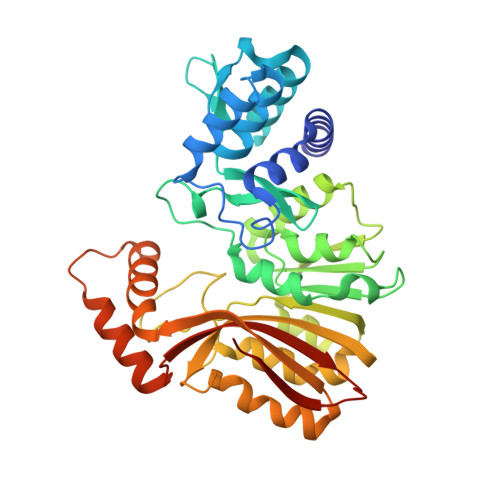
Structure and mechanism for iterative amide N -methylation in the biosynthesis of channel-forming peptide cytotoxins.
Cogan, D.P., Bhushan, A., Reyes, R., Zhu, L., Piel, J., Nair, S.K.(2022) Proc Natl Acad Sci U S A 119: e2116578119-e2116578119
- PubMed: 35316135 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2116578119
- Primary Citation Related Structures:
7RC2, 7RC3, 7RC4, 7RC5, 7RC6 - PubMed Abstract:
SignificanceThe channel-forming proteusins are bacterial helical peptides that allow permeation of positively charged ions to influence membrane potential and cellular physiology. We biochemically characterize the effect of two critical posttranslational modifications on the secondary structure of the peptide substrate. We determine how a methyl group can be added to the side chains of D-Asn residues in a peptide substrate and show how flanking residues influence selectivity. These studies should foster the development of small-molecule peptide ion channels as therapeutics.
- Department of Biochemistry, University of Illinois at Urbana-Champaign, Urbana, IL 61801.
Organizational Affiliation: