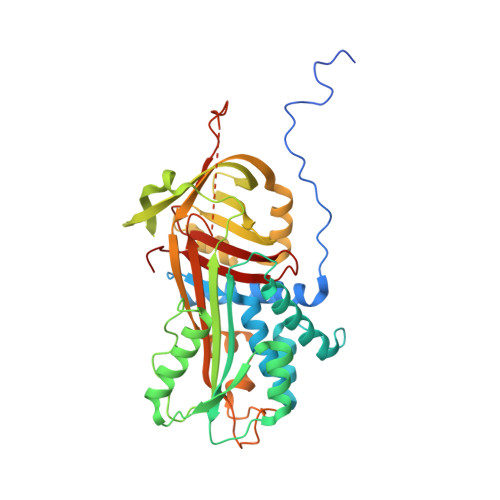
Structural and Functional Characterization of a Biliverdin-Binding Near-Infrared Fluorescent Protein From the Serpin Superfamily.
Manoilov, K.Y., Ghosh, A., Almo, S.C., Verkhusha, V.V.(2021) J Mol Biology 434: 167359-167359
- PubMed: 34798132 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2021.167359
- Primary Citation Related Structures:
7RBW - PubMed Abstract:
Biliverdin-binding serpins (BBSs) are proteins that are responsible for coloration in amphibians and fluoresce in the near-infrared (NIR) spectral region. Here we produced the first functional recombinant BBS of the polka-dot treefrog Boana punctata (BpBBS), assembled with its biliverdin (BV) chromophore, and report its biochemical and photochemical characterization. We determined the crystal structure of BpBBS at 2.05 Å resolution, which demonstrated its structural homology to the mammalian protease inhibitor alpha-1-antitrypsin. BV interaction with BpBBS was studied and it was found that the N-terminal polypeptide (residues 19-50) plays a critical role in the BV binding. By comparing BpBBS with the available NIR fluorescent proteins and expressing it in mammalian cells, we demonstrated its potential as a NIR imaging probe. These results provide insight into the non-inhibitory function of serpins, provide a basis for improving their performance in mammalian cells, and suggest possible paths for the development of BBS-based fluorescent probes.
- Department of Anatomy and Structural Biology, Albert Einstein College of Medicine, Bronx, NY 10461, USA.
Organizational Affiliation: