Molecular basis of differential HLA class I-restricted T cell recognition of a highly networked HIV peptide.
Li, X., Singh, N.K., Collins, D.R., Ng, R., Zhang, A., Lamothe-Molina, P.A., Shahinian, P., Xu, S., Tan, K., Piechocka-Trocha, A., Urbach, J.M., Weber, J.K., Gaiha, G.D., Takou Mbah, O.C., Huynh, T., Cheever, S., Chen, J., Birnbaum, M., Zhou, R., Walker, B.D., Wang, J.H.(2023) Nat Commun 14: 2929-2929
- PubMed: 37217466 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38573-8
- Primary Citation Related Structures:
7R7V, 7R7W, 7R7X, 7R7Y, 7R7Z, 7R80 - PubMed Abstract:
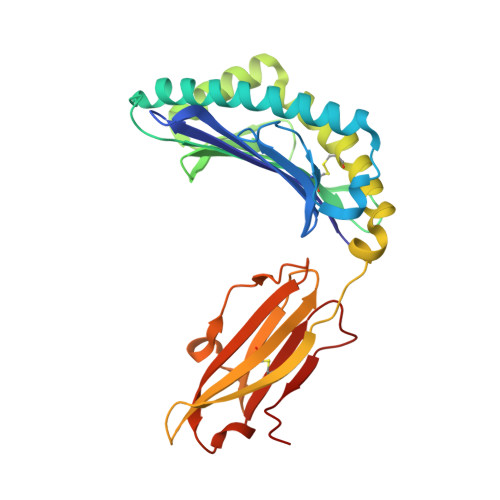
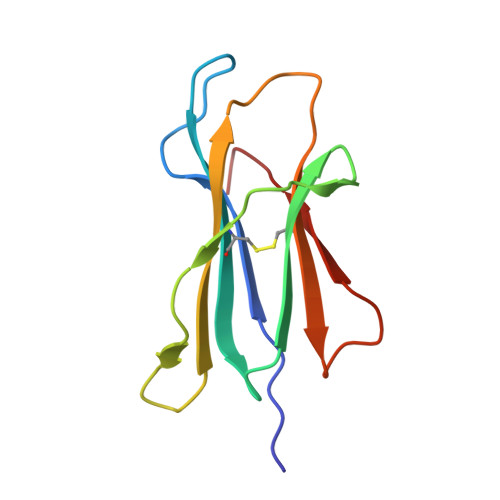
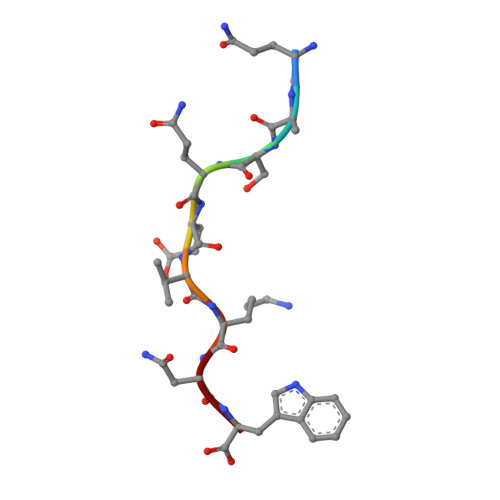
Cytotoxic-T-lymphocyte (CTL) mediated control of HIV-1 is enhanced by targeting highly networked epitopes in complex with human-leukocyte-antigen-class-I (HLA-I). However, the extent to which the presenting HLA allele contributes to this process is unknown. Here we examine the CTL response to QW9, a highly networked epitope presented by the disease-protective HLA-B57 and disease-neutral HLA-B53. Despite robust targeting of QW9 in persons expressing either allele, T cell receptor (TCR) cross-recognition of the naturally occurring variant QW9_S3T is consistently reduced when presented by HLA-B53 but not by HLA-B57. Crystal structures show substantial conformational changes from QW9-HLA to QW9_S3T-HLA by both alleles. The TCR-QW9-B53 ternary complex structure manifests how the QW9-B53 can elicit effective CTLs and suggests sterically hindered cross-recognition by QW9_S3T-B53. We observe populations of cross-reactive TCRs for B57, but not B53 and also find greater peptide-HLA stability for B57 in comparison to B53. These data demonstrate differential impacts of HLAs on TCR cross-recognition and antigen presentation of a naturally arising variant, with important implications for vaccine design.
- School of Life Sciences, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, Anhui, 230027, China. xiaolong@crystal.harvard.edu.
Organizational Affiliation: