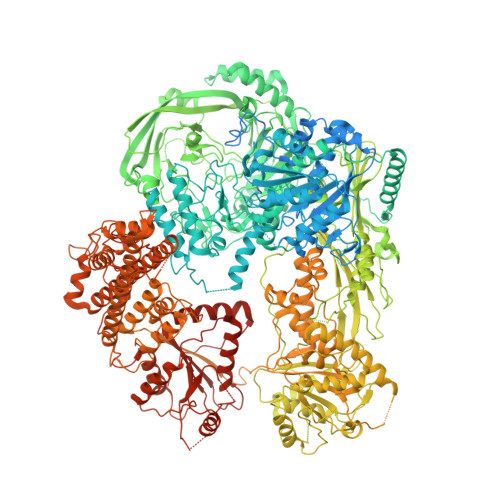
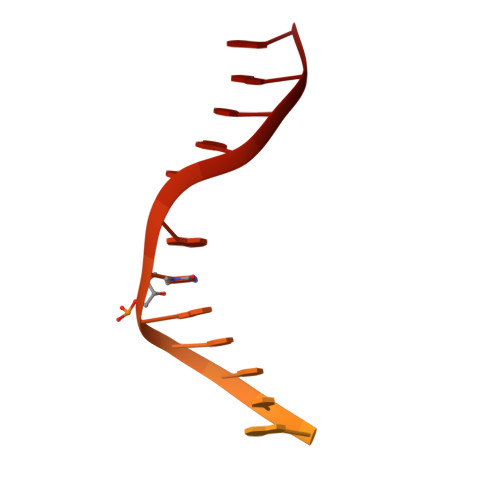
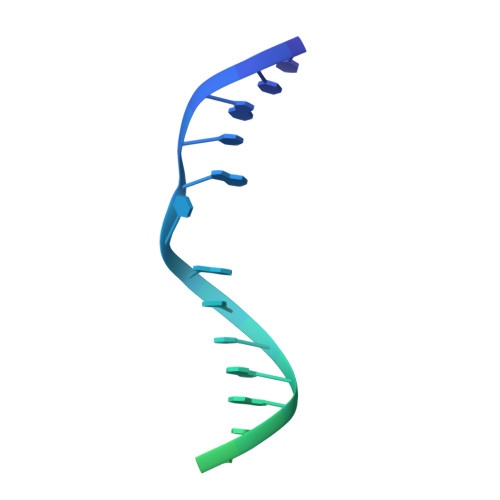
Structural insights into DNMT5-mediated ATP-dependent high-fidelity epigenome maintenance.
Wang, J., Catania, S., Wang, C., de la Cruz, M.J., Rao, B., Madhani, H.D., Patel, D.J.(2022) Mol Cell 82: 1186-1198.e6
- PubMed: 35202575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2022.01.028
- Primary Citation Related Structures:
7R76, 7R77, 7R78, 7T02 - PubMed Abstract:
Epigenetic evolution occurs over million-year timescales in Cryptococcus neoformans and is mediated by DNMT5, the first maintenance type cytosine methyltransferase identified in the fungal or protist kingdoms, the first dependent on adenosine triphosphate (ATP), and the most hemimethyl-DNA-specific enzyme known. To understand these novel properties, we solved cryo-EM structures of CnDNMT5 in three states. These studies reveal an elaborate allosteric cascade in which hemimethylated DNA binding first activates the SNF2 ATPase domain by a large rigid body rotation while the target cytosine partially flips out of the DNA duplex. ATP binding then triggers striking structural reconfigurations of the methyltransferase catalytic pocket to enable cofactor binding, completion of base flipping, and catalysis. Bound unmethylated DNA does not open the catalytic pocket and is instead ejected upon ATP binding, driving high fidelity. This unprecedented chaperone-like, enzyme-remodeling role of the SNF2 ATPase domain illuminates how energy is used to enable faithful epigenetic memory.
- Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA. Electronic address: wangj1@mskcc.org.
Organizational Affiliation: