Halo complexes of gold(I) containing glycoconjugate carbene ligands: synthesis, characterization, cytotoxicity and interaction with proteins and DNA model systems.
Annunziata, A., Ferraro, G., Cucciolito, M.E., Imbimbo, P., Tuzi, A., Monti, D.M., Merlino, A., Ruffo, F.(2022) Dalton Trans 51: 10475-10485
- PubMed: 35766118 Search on PubMed
- DOI: https://doi.org/10.1039/d2dt00423b
- Primary Citation Related Structures:
7R1P, 7R1Q - PubMed Abstract:
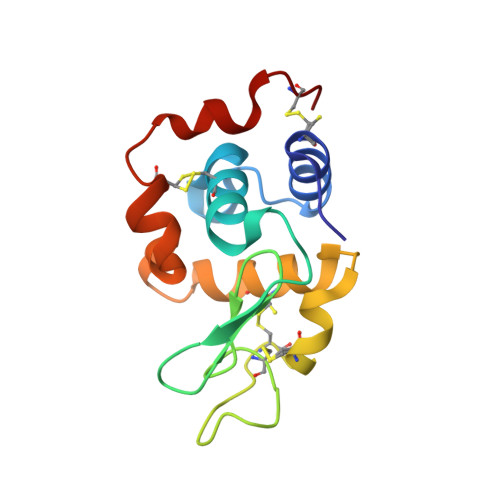
New neutral carbene complexes of gold(I) [Au(Im-Me)X] (X = Cl, Au1; X = Br, Au2; X = I, Au3) have been synthesized and fully characterized by different techniques, including NMR and UV-vis absorption spectroscopy and single crystal X-ray diffraction. The carbene ligand Im-Me is decorated with a glucoside fragment via a triazole linker, obtainable through a click chemistry reaction. The compounds retain the Au-NHC fragment in aqueous solvents, and an equilibrium between the neutral halo- and the cationic di-carbene form [Au(Im-Me) 2 ] + is observed, whose extent follows the trend Au1 < Au2 < Au3. Cytotoxicity studies on two cancer and two non-tumorigenic cell lines reflect the solution behavior, as a certain difference among the complexes was disclosed, with the iodo complex Au3 being more active and selective. The compounds interact with both DNA and protein model systems. The X-ray structure of the adduct formed upon the reaction of Au1 with bovine pancreatic ribonuclease (RNase A) reveals Au binding at the side chain of His105 of both protein molecules A and B of the asymmetric unit. The binding of gold atoms at both the nitrogen atoms of the imidazole ring of His15 and at the N-terminal tail has been found in the adduct formed with hen egg white lysozyme.
- Dipartimento di Scienze Chimiche, Università di Napoli Federico II, Complesso Universitario di Monte Sant'Angelo, Via Cintia 21, 80126, Napoli, Italy. dariamaria.monti@unina.it.
Organizational Affiliation: