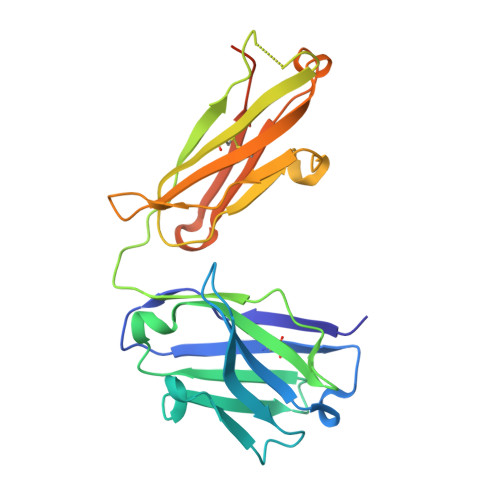
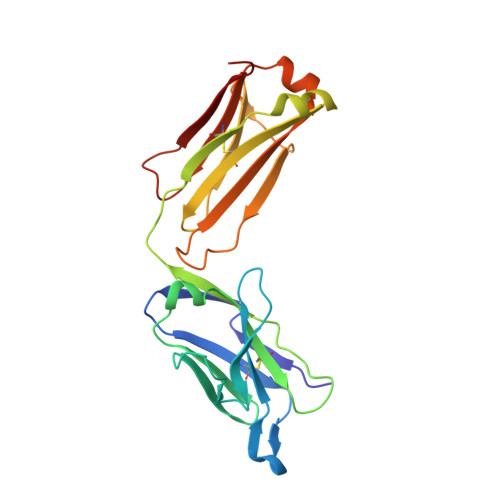
Antibody Recognition of Different Staphylococcus aureus Wall Teichoic Acid Glycoforms.
Di Carluccio, C., Soriano-Maldonado, P., Berni, F., de Haas, C.J.C., Temming, A.R., Hendriks, A., Ali, S., Molinaro, A., Silipo, A., van Sorge, N.M., van Raaij, M.J., Codee, J.D.C., Marchetti, R.(2022) ACS Cent Sci 8: 1383-1392
- PubMed: 36313161 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acscentsci.2c00125
- Primary Citation Related Structures:
7QXC, 7QXD, 7QXE - PubMed Abstract:
Wall teichoic acids (WTAs) are glycopolymers decorating the surface of Gram-positive bacteria and potential targets for antibody-mediated treatments against Staphylococcus aureus , including methicillin-resistant (MRSA) strains. Through a combination of glycan microarray, synthetic chemistry, crystallography, NMR, and computational studies, we unraveled the molecular and structural details of fully defined synthetic WTA fragments recognized by previously described monoclonal antibodies (mAbs 4461 and 4497). Our results unveiled the structural requirements for the discriminatory recognition of α- and β-GlcNAc-modified WTA glycoforms by the complementarity-determining regions (CDRs) of the heavy and light chains of the mAbs. Both mAbs interacted not only with the sugar moiety but also with the phosphate groups as well as residues in the ribitol phosphate (RboP) units of the WTA backbone, highlighting their significant role in ligand specificity. Using elongated WTA fragments, containing two sugar modifications, we also demonstrated that the internal carbohydrate moiety of α-GlcNAc-modified WTA is preferentially accommodated in the binding pocket of mAb 4461 with respect to the terminal moiety. Our results also explained the recently documented cross-reactivity of mAb 4497 for β-1,3/β-1,4-GlcNAc-modified WTA, revealing that the flexibility of the RboP backbone is crucial to allow positioning of both glycans in the antibody binding pocket.
- Department of Chemical Sciences, University of Naples Federico II, Via Cinthia 4, 80126Naples, Italy.
Organizational Affiliation: