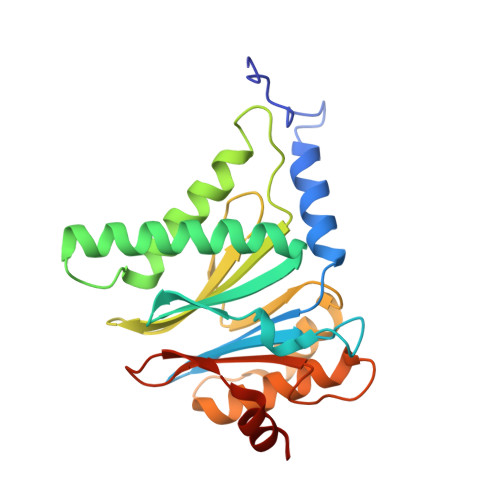
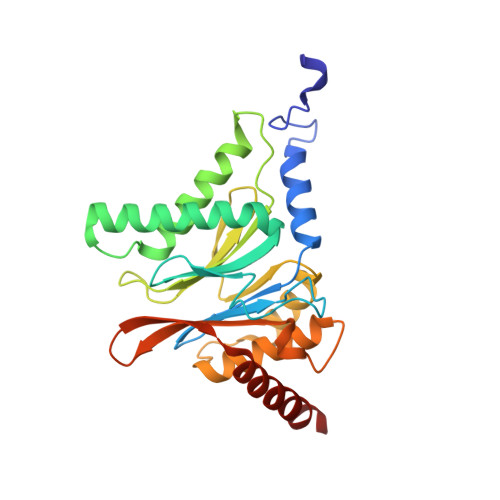
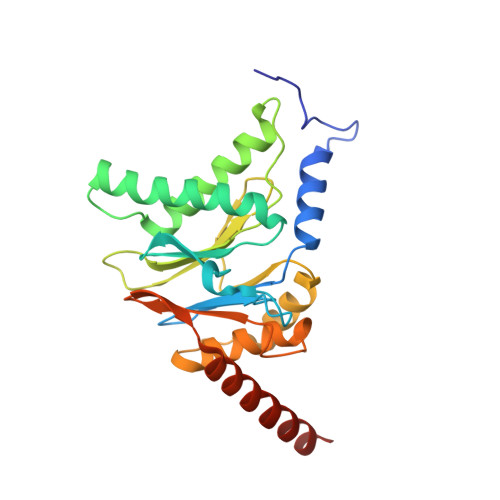
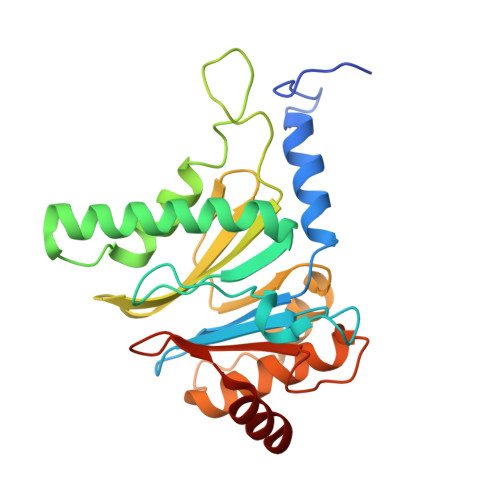
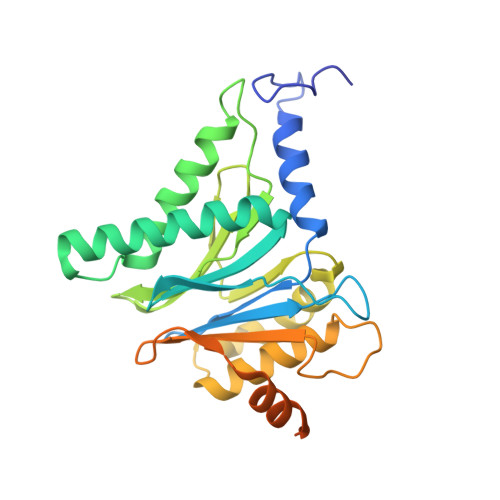
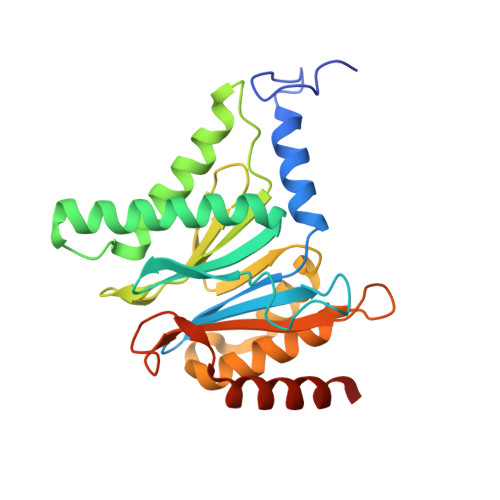
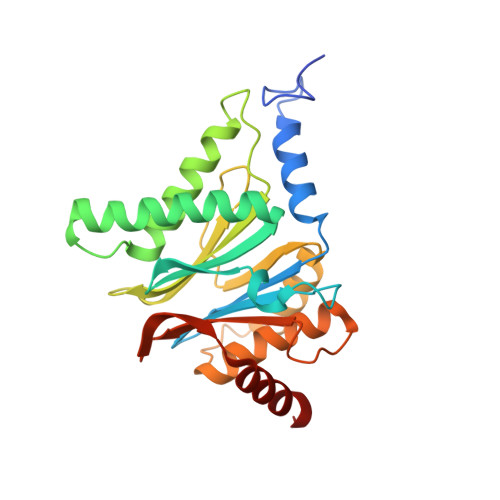
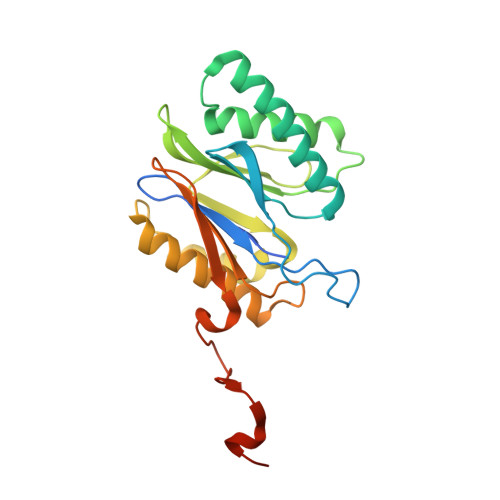
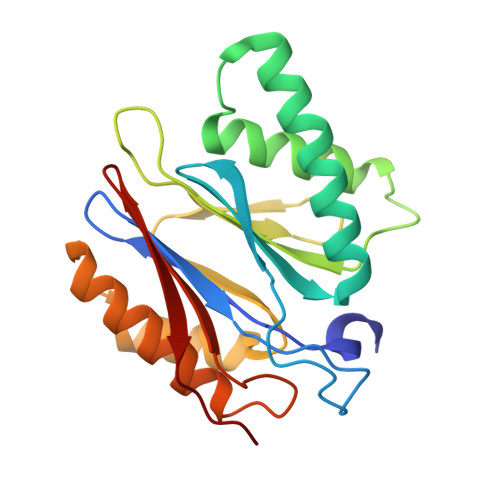
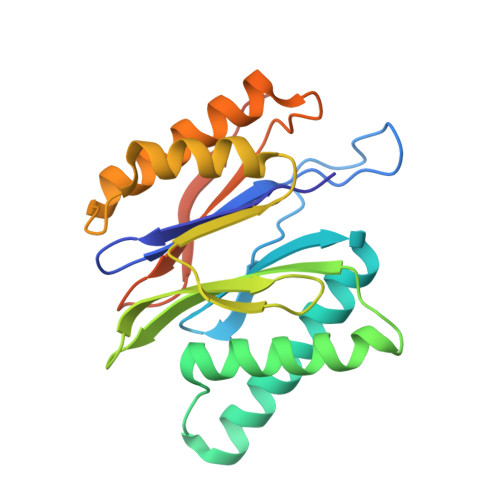
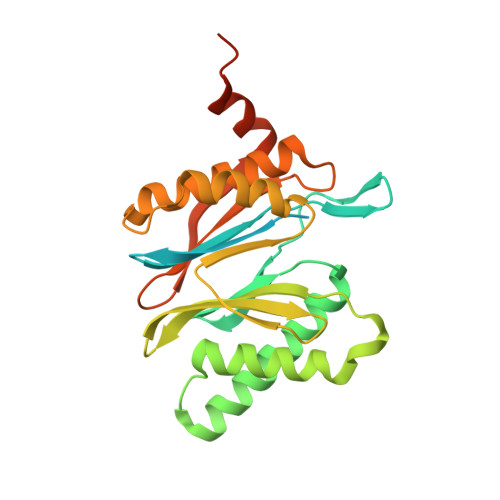
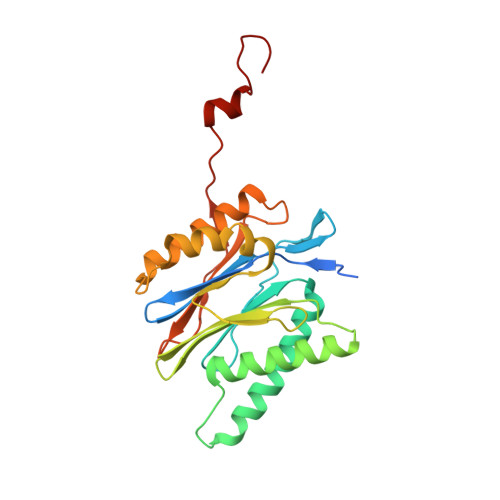
Cryo-EM structure of the plant 26S proteasome.
Kandolf, S., Grishkovskaya, I., Belacic, K., Bolhuis, D.L., Amann, S., Foster, B., Imre, R., Mechtler, K., Schleiffer, A., Tagare, H.D., Zhong, E.D., Meinhart, A., Brown, N.G., Haselbach, D.(2022) Plant Commun 3: 100310-100310
- PubMed: 35576154 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.xplc.2022.100310
- Primary Citation Related Structures:
7QVE - PubMed Abstract:
Targeted proteolysis is a hallmark of life. It is especially important in long-lived cells that can be found in higher eukaryotes, like plants. This task is mainly fulfilled by the ubiquitin-proteasome system. Thus, proteolysis by the 26S proteasome is vital to development, immunity, and cell division. Although the yeast and animal proteasomes are well characterized, there is only limited information on the plant proteasome. We determined the first plant 26S proteasome structure from Spinacia oleracea by single-particle electron cryogenic microscopy at an overall resolution of 3.3 Å. We found an almost identical overall architecture of the spinach proteasome compared with the known structures from mammals and yeast. Nevertheless, we noticed a structural difference in the proteolytic active β1 subunit. Furthermore, we uncovered an unseen compression state by characterizing the proteasome's conformational landscape. We suspect that this new conformation of the 20S core protease, in correlation with a partial opening of the unoccupied gate, may contribute to peptide release after proteolysis. Our data provide a structural basis for the plant proteasome, which is crucial for further studies.
- Research Institute of Molecular Pathology (IMP), Vienna BioCenter (VBC), Campus-Vienna-BioCenter 1, 1030 Vienna, Austria.
Organizational Affiliation: